| [Ref.: #34205] |
Culture collection no. |
CIP 78.34, ATCC 19113, CCM 5578, NCTC 5105, JCM 7673, SLCC 2373 |
| [Ref.: #92626] |
 SI-ID 36856 SI-ID 36856
|
* |
 |
Literature: |
Only first 10 entries are displayed. Click here to see all.Click here to see only first 10 entries. |
|
Topic |
Title |
Authors |
Journal |
DOI |
Year |
|
| Biotechnology |
Antibacterial effects of hydrogen peroxide and caprylic acid on selected foodborne bacteria. |
Vyrostkova J, Pipova M, Semjon B, Jevinova P, Regecova I, Malova J |
Pol J Vet Sci |
10.24425/pjvs.2020.134689 |
2020 |
* |
| Biotechnology |
Detection of sublethal thermal injury in Salmonella enterica serotype typhimurium and Listeria monocytogenes using Fourier transform infrared (FT-IR) spectroscopy (4000 to 600 cm(-1)). |
Al-Qadiri HM, Lin M, Al-Holy MA, Cavinato AG, Rasco BA |
J Food Sci |
10.1111/j.1750-3841.2007.00640.x |
2008 |
* |
| Pathogenicity |
Bactericidal effect of gentamicin-induced membrane vesicles derived from Pseudomonas aeruginosa PAO1 on gram-positive bacteria. |
MacDonald KL, Beveridge TJ |
Can J Microbiol |
10.1139/w02-077 |
2002 |
* |
| Pathogenicity |
Resistance of Listeria monocytogenes to the bacteriocin nisin. |
Davies EA, Adams MR |
Int J Food Microbiol |
10.1016/0168-1605(94)90064-7 |
1994 |
* |
| Phylogeny |
Determination of virulence of different strains of Listeria monocytogenes and Listeria innocua by oral inoculation of pregnant mice. |
Lammerding AM, Glass KA, Gendron-Fitzpatrick A, Doyle MP |
Appl Environ Microbiol |
10.1128/aem.58.12.3991-4000.1992 |
1992 |
* |
| Pathogenicity |
Cytopathogenic effects in enterocytelike Caco-2 cells differentiate virulent from avirulent Listeria strains. |
Pine L, Kathariou S, Quinn F, George V, Wenger JD, Weaver RE |
J Clin Microbiol |
10.1128/jcm.29.5.990-996.1991 |
1991 |
* |
|
Suppurative gastritis in BALB/c mice infected with Listeria monocytogenes via the intragastric route. |
Park JH, Park YH, Seok SH, Cho SA, Kim DJ, Lee HY, Kim SH, Park JH |
J Comp Pathol |
10.1016/j.jcpa.2003.10.001 |
2004 |
* |
|
Low temperature virulence of Listeria monocytogenes in the avian embryo. |
Wood LV, Woodbine M |
Zentralbl Bakteriol Orig A |
|
1979 |
* |
|
 References References-
-
| #34205 |
Collection of Institut Pasteur ; Curators of the CIP;
CIP 78.34
|
-
| #67770 |
Japan Collection of Microorganism (JCM) ; Curators of the JCM;
|
-
| #68371 |
Automatically annotated from API 50CH acid .
|
-
| #68382 |
Automatically annotated from API zym .
|
-
| #69479 |
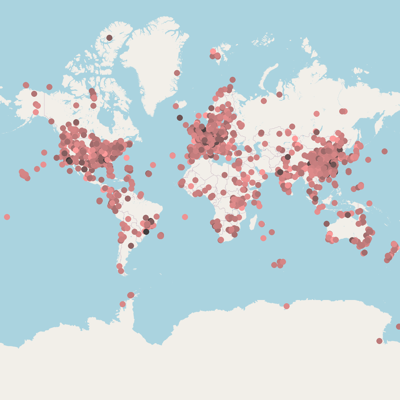
João F Matias Rodrigues, Janko Tackmann,Gregor Rot, Thomas SB Schmidt, Lukas Malfertheiner, Mihai Danaila,Marija Dmitrijeva, Daniela Gaio, Nicolas Näpflin and Christian von Mering. University of Zurich.:
MicrobeAtlas 1.0 beta
.
|
-
| #92626 |
Reimer, L.C., Lissin, A.,Schober, I., Witte,J.F., Podstawka, A., Lüken, H., Bunk, B.,Overmann, J.:
StrainInfo: A central database for resolving microbial strain identifiers
.
(
DOI 10.60712/SI-ID36856.1 )
|
- * These data were automatically processed and therefore are not curated
Change proposal
Successfully sent
|
Name and taxonomic classification

Morphology

Culture and growth conditions

Physiology and metabolism

Isolation, sampling and environmental information

Safety information

Sequence information

External links

References