-
Availability in culture collections
 External links External links
| [Ref.: #9300] |
Culture collection no. |
DSM 40088, AS 4.1370, ATCC 19793, CBS 545.68, ETH 28497, IFO 12804, ISP 5088, JCM 4251, JCM 4599, KCC S-0251, KCC S-0599, NBRC 12804, NCIB 9219, NRRL 2466, RIA 1072, BCRC 11514, CCUG 11108, CGMCC 4.1370, IFM 1181, IMET 43503, LMG 19395, LMG 5980, MTCC 2525, NCIMB 9219 |
| [Ref.: #85485] |
 SI-ID 13856 SI-ID 13856
|
* |
 |
Literature: |
|
Topic |
Title |
Authors |
Journal |
DOI |
Year |
|
| Phylogeny |
The search for synonyms among streptomycetes by using SDS-PAGE of whole-cell proteins. Emendation of the species Streptomyces aurantiacus, Streptomyces cacaoi subsp. cacaoi, Streptomyces caeruleus and Streptomyces violaceus. |
Lanoot B, Vancanneyt M, Cleenwerck I, Wang L, Li W, Liu Z, Swings J |
Int J Syst Evol Microbiol |
10.1099/00207713-52-3-823 |
2002 |
* |
| Phylogeny |
Streptomyces corynorhini sp. nov., isolated from Townsend's big-eared bats (Corynorhinus townsendii). |
Hamm PS, Caimi NA, Northup DE, Valdez EW, Buecher DC, Dunlap CA, Labeda DP, Porras-Alfaro A |
Antonie Van Leeuwenhoek |
10.1007/s10482-019-01261-z |
2019 |
* |
| Phylogeny |
Streptomyces camponoticapitis sp. nov., an actinomycete isolated from the head of an ant (Camponotus japonicus Mayr). |
Li Y, Ye L, Wang X, Zhao J, Ma Z, Yan K, Xiang W, Liu C |
Int J Syst Evol Microbiol |
10.1099/ijsem.0.001276 |
2016 |
* |
| Metabolism |
High frequency transformation of Streptomyces niveus protoplasts by plasmid DNA. |
Hussain HA, Ritchie DA |
J Appl Bacteriol |
10.1111/j.1365-2672.1991.tb03811.x |
1991 |
* |
| Enzymology |
Genetic instability in Streptomyces niveus plasmid pSN2: in vivo formation of deletion derivatives. |
Hussain HA, Mitchell JI, Ritchie DA |
Arch Microbiol |
10.1007/BF00245235 |
1990 |
* |
|
 References References-
| #9300 |
Leibniz Institut DSMZ-Deutsche Sammlung von Mikroorganismen und Zellkulturen GmbH ; Curators of the DSMZ;
DSM 40088
|
-
-
-
-
-
| #67770 |
Japan Collection of Microorganism (JCM) ; Curators of the JCM;
|
-
-
-
| #85485 |
Reimer, L.C., Lissin, A.,Schober, I., Witte,J.F., Podstawka, A., Lüken, H., Bunk, B.,Overmann, J.:
StrainInfo: A central database for resolving microbial strain identifiers
.
(
DOI 10.60712/SI-ID13856.1 )
|
- * These data were automatically processed and therefore are not curated
Change proposal
Successfully sent
|
|
Name and taxonomic classification

Morphology

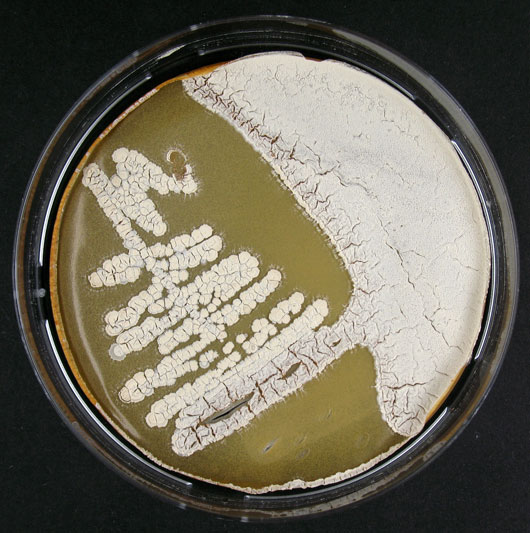
Culture and growth conditions

Physiology and metabolism

Isolation, sampling and environmental information

Safety information

Sequence information

Genome-based predictions

External links

References





