| [Ref.: #1402] |
Sample type/isolated from |
alcohol turned to vinegar |
| |
| [Ref.: #46757] |
Sample type/isolated from |
Beech-wood shavings of vinegar plant |
| [Ref.: #46757] |
Sampling date |
1922 |
| |
| [Ref.: #67770] |
Sample type/isolated from |
Alcohol turned to vinegar |
| |
| [Ref.: #115989] |
Sample type/isolated from |
Food, Alcohol vinegar |
|
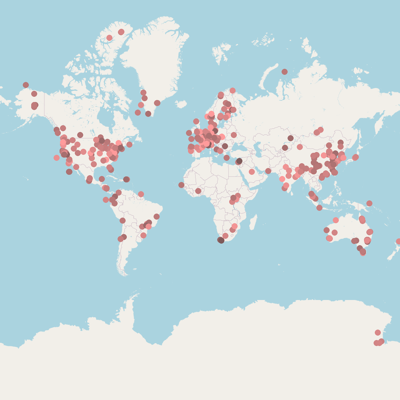
* marker position based on {}
|
|
Isolation sources categories |
| #Engineered |
#Food production |
- |
| #Environmental |
#Microbial community |
- |
|
 |
-
Availability in culture collections
 External links External links
| [Ref.: #1402] |
Culture collection no. |
DSM 3508, ATCC 15973, NCIB 8621, CCUG 18122, LMG 1261-t1, LMG 1261 1, CCTM 3043, NBRC 14818, JCM 7641, BCRC 11688, CCM 3620, CECT 298, CGMCC 1.1809, CIP 103111, ICMP 8807, IFO 14818, LMD 23.1, LMG 1261, LMG 1504, NCCB 23001, NCIMB 8621, VTT E-042567, VTT E-97828 |
| [Ref.: #69687] |
 SI-ID 92754 SI-ID 92754
|
* |
 |
Literature: |
Only first 10 entries are displayed. Click here to see all.Click here to see only first 10 entries. |
|
Topic |
Title |
Authors |
Journal |
DOI |
Year |
|
| Genetics |
Complete Genome Sequence of the Acetic Acid Bacterium Acetobacter aceti NBRC 14818. |
Arai H, Kameya M, Ishii M |
Microbiol Resour Announc |
10.1128/MRA.01039-20 |
2020 |
* |
| Phylogeny |
Acetobacter musti sp. nov., isolated from Bobal grape must. |
Ferrer S, Manes-Lazaro R, Benavent-Gil Y, Yepez A, Pardo I |
Int J Syst Evol Microbiol |
10.1099/ijsem.0.000818 |
2015 |
* |
| Metabolism |
Role of the glyoxylate pathway in acetic acid production by Acetobacter aceti. |
Sakurai K, Yamazaki S, Ishii M, Igarashi Y, Arai H |
J Biosci Bioeng |
10.1016/j.jbiosc.2012.07.017 |
2012 |
* |
| Metabolism |
Changes in the gene expression profile of Acetobacter aceti during growth on ethanol. |
Sakurai K, Arai H, Ishii M, Igarashi Y |
J Biosci Bioeng |
10.1016/j.jbiosc.2011.11.005 |
2011 |
* |
| Metabolism |
Transcriptome response to different carbon sources in Acetobacter aceti. |
Sakurai K, Arai H, Ishii M, Igarashi Y |
Microbiology (Reading) |
10.1099/mic.0.045906-0 |
2010 |
* |
| Enzymology |
IS1032 from Acetobacter xylinum, a new mobile insertion sequence. |
Iversen T, Standal R, Pedersen T, Coucheron DH |
Plasmid |
10.1006/plas.1994.1043 |
1994 |
* |
| Phylogeny |
Numerical Analysis of Phenotypic Features and Protein Gel Electrophoregrams of a Wide Variety of Acetobacter strains. Proposal for the Improvement of the Taxonomy of the Genus Acetobacter Beijerinck 1898, 215. |
Gossele F, Swings J, Kersters K, Pauwels P, De Ley J |
Syst Appl Microbiol |
10.1016/S0723-2020(83)80020-4 |
1983 |
* |
|
 References References-
| #1402 |
Leibniz Institut DSMZ-Deutsche Sammlung von Mikroorganismen und Zellkulturen GmbH ; Curators of the DSMZ;
DSM 3508
|
-
-
-
| #41703 |
; Curators of the CIP;
|
-
| #46757 |
Culture Collection University of Gothenburg (CCUG) ; Curators of the CCUG;
CCUG 18122
|
-
-
| #66794 |
Antje Chang, Lisa Jeske, Sandra Ulbrich, Julia Hofmann, Julia Koblitz, Ida Schomburg, Meina Neumann-Schaal, Dieter Jahn, Dietmar Schomburg:
BRENDA, the ELIXIR core data resource in 2021: new developments and updates.
Nucleic Acids Res. 49:
D498 - D508
2020 (
DOI 10.1093/nar/gkaa1025 , PubMed 33211880 )
|
-
| #67770 |
Japan Collection of Microorganism (JCM) ; Curators of the JCM;
|
-
| #68368 |
Automatically annotated from API 20E .
|
-
| #68382 |
Automatically annotated from API zym .
|
-
| #69479 |
João F Matias Rodrigues, Janko Tackmann,Gregor Rot, Thomas SB Schmidt, Lukas Malfertheiner, Mihai Danaila,Marija Dmitrijeva, Daniela Gaio, Nicolas Näpflin and Christian von Mering. University of Zurich.:
MicrobeAtlas 1.0 beta
.
|
-
-
-
| #69687 |
Reimer, L.C., Lissin, A.,Schober, I., Witte,J.F., Podstawka, A., Lüken, H., Bunk, B.,Overmann, J.:
StrainInfo: A central database for resolving microbial strain identifiers
.
(
DOI 10.60712/SI-ID92754.1 )
|
-
| #115989 |
Collection of Institut Pasteur ; Curators of the CIP;
CIP 103111
|
- * These data were automatically processed and therefore are not curated
Change proposal
Successfully sent
|
|
Name and taxonomic classification

Morphology

Culture and growth conditions

Physiology and metabolism

Isolation, sampling and environmental information

Safety information

Sequence information

Genome-based predictions

External links

References