| [Ref.: #35278] |
Culture collection no. |
CIP 102998, IAM 1802, IFO 3281, JCM 20268, LMG 1512, NBRC 3281 |
| [Ref.: #93241] |
 SI-ID 14539 SI-ID 14539
|
* |
 |
Literature: |
|
Topic |
Title |
Authors |
Journal |
DOI |
Year |
|
| Biotechnology |
Production of D-lyxose from D-glucose by microbial and enzymatic reactions. |
Ahmed Z, Sasahara H, Bhuiyan SH, Saiki T, Shimonishi T, Takada G, Izumori K |
J Biosci Bioeng |
10.1016/s1389-1723(00)87100-5 |
1999 |
* |
| Metabolism |
Growth characteristics and oxidative capacity of Acetobacter aceti IFO 3281: implications for L-ribulose production. |
Kylma AK, Granstrom T, Leisola M |
Appl Microbiol Biotechnol |
10.1007/s00253-003-1406-4 |
2003 |
* |
| Enzymology |
Purification of restriction endonuclease from Acetobacter aceti IFO 3281 (AatII) and its properties. |
Sato H, Suzuki T, Yamada Y |
Agric Biol Chem |
|
1990 |
* |
| Enzymology |
New restriction endonucleases from Acetobacter aceti and Bacillus aneurinolyticus. |
Sugisaki H, Maekawa Y, Kanazawa S, Takanami M |
Nucleic Acids Res |
10.1093/nar/10.19.5747 |
1982 |
* |
|
Biochemical preparation of L-ribose and L-arabinose from ribitol: a new approach. |
Ahmed Z, Shimonishi T, Bhuiyan SH, Utamura M, Takada G, Izumori K |
J Biosci Bioeng |
10.1016/s1389-1723(99)80225-4 |
1999 |
* |
|
 References References-
-
| #35278 |
Collection of Institut Pasteur ; Curators of the CIP;
CIP 102998
|
-
-
| #67770 |
Japan Collection of Microorganism (JCM) ; Curators of the JCM;
|
-
| #69479 |
João F Matias Rodrigues, Janko Tackmann,Gregor Rot, Thomas SB Schmidt, Lukas Malfertheiner, Mihai Danaila,Marija Dmitrijeva, Daniela Gaio, Nicolas Näpflin and Christian von Mering. University of Zurich.:
MicrobeAtlas 1.0 beta
.
|
-
-
-
| #93241 |
Reimer, L.C., Lissin, A.,Schober, I., Witte,J.F., Podstawka, A., Lüken, H., Bunk, B.,Overmann, J.:
StrainInfo: A central database for resolving microbial strain identifiers
.
(
DOI 10.60712/SI-ID14539.1 )
|
- * These data were automatically processed and therefore are not curated
Change proposal
Successfully sent
|
Name and taxonomic classification

Morphology

Culture and growth conditions

Physiology and metabolism

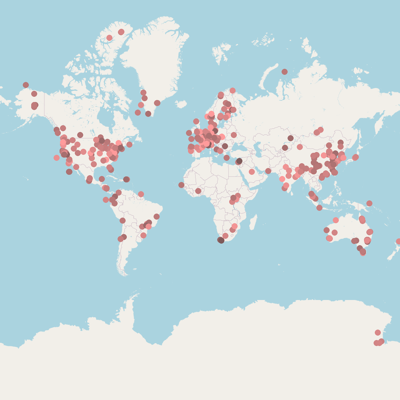
Isolation, sampling and environmental information

Safety information

Sequence information

Genome-based predictions

External links

References