| [Ref.: #19349] |
Sample type/isolated from |
root of soybean (Glycine max (L.) Merr) |
| [Ref.: #19349] |
Host species |
Glycine max |
| [Ref.: #19349] |
Geographic location (country and/or sea, region) |
Heilongjiang province, Hulin |
| [Ref.: #19349] |
Country |
China |
| [Ref.: #19349] |
Country ISO 3 Code |
CHN |
| [Ref.: #19349] |
Continent |
Asia |
|
* marker position based on {}
|
|
Isolation sources categories |
| #Host |
#Plants |
#Herbaceous plants (Grass,Crops) |
| #Host Body-Site |
#Plant |
#Root (Rhizome) |
|
 |
-
Availability in culture collections
 External links External links
| [Ref.: #19349] |
Culture collection no. |
DSM 45728, BCC 72029, CGMCC 4.7036 |
 |
Literature: |
|
Topic |
Title |
Authors |
Journal |
DOI |
Year |
|
| Phylogeny |
Actinoplanes flavus sp. nov., a novel cellulase-producing actinobacterium isolated from coconut palm rhizosphere soil. |
Luo X, Sun X, Huang Z, He C, Zhao J, Xiang W, Song J, Wang X |
Int J Syst Evol Microbiol |
10.1099/ijsem.0.004990 |
2021 |
* |
| Phylogeny |
Actinoplanes hulinensis sp. nov., a novel actinomycete isolated from soybean root (Glycine max (L.) Merr). |
Shen Y, Liu C, Wang X, Zhao J, Jia F, Zhang Y, Wang L, Yang D, Xiang W |
Antonie Van Leeuwenhoek |
10.1007/s10482-012-9809-9 |
2012 |
* |
| Phylogeny |
Actinoplanes subglobosus sp. nov., isolated from mixed deciduous forest soil. |
Ngaemthao W, Chunhametha S, Suriyachadkun C |
Int J Syst Evol Microbiol |
10.1099/ijsem.0.001440 |
2016 |
* |
|
 References References-
| #19349 |
Leibniz Institut DSMZ-Deutsche Sammlung von Mikroorganismen und Zellkulturen GmbH ; Curators of the DSMZ;
DSM 45728
|
-
-
-
-
- * These data were automatically processed and therefore are not curated
Change proposal
Successfully sent
|
|
Name and taxonomic classification

Morphology

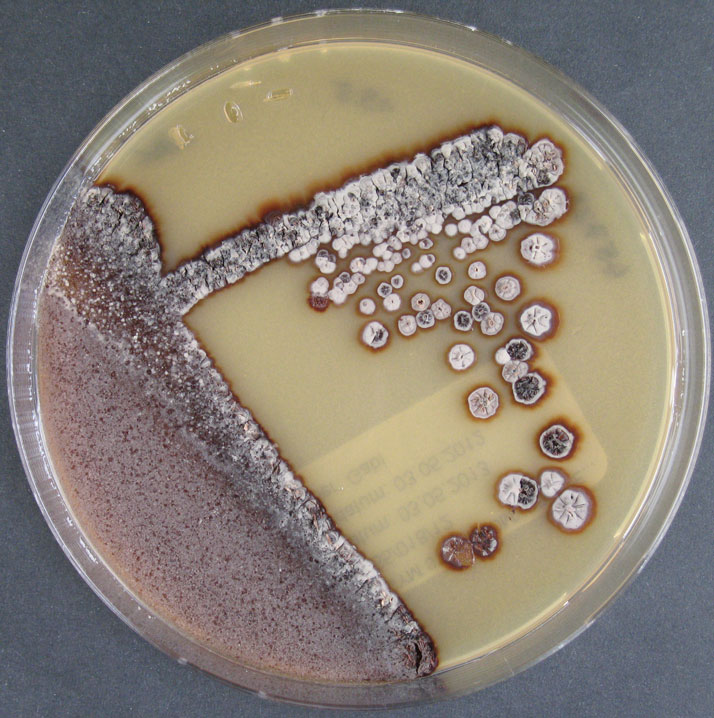
Culture and growth conditions

Physiology and metabolism

Isolation, sampling and environmental information

Sequence information

Genome-based predictions

External links

References



