| [Ref.: #9245] |
Sample type/isolated from |
soil |
|
* marker position based on {}
|
|
Isolation sources categories |
| #Environmental |
#Terrestrial |
#Soil |
|
 |
-
Information on genomic background e.g. entries in nucleic sequence databass
 Sequence information Sequence information
|
16S Sequence information: |
Only first 5 entries are displayed. Click here to see all.Click here to see only first 5 entries. |
|
|
Sequence accession description |
Seq. accession number |
Sequence length (bp) |
Sequence database |
Associated NCBI tax ID |
|
| [Ref.: #20218] |
Streptomyces sp. NTG2 gene for 16S ribosomal RNA, partial sequence |
AB920573 |
1252 |
|
1571709 tax ID tax ID |
* |
| [Ref.: #20218] |
Streptomyces sp. NTG5 gene for 16S ribosomal RNA, partial sequence |
AB920576 |
1286 |
|
1571712 tax ID tax ID |
* |
| [Ref.: #20218] |
Streptomyces sp. NTGn11 gene for 16S ribosomal RNA, partial sequence |
AB920578 |
1303 |
|
1571714 tax ID tax ID |
* |
| [Ref.: #20218] |
Streptomyces sp. NTHn1 gene for 16S ribosomal RNA, partial sequence |
AB920579 |
1306 |
|
1571715 tax ID tax ID |
* |
| [Ref.: #20218] |
Streptomyces sp. NTHn2 gene for 16S ribosomal RNA, partial sequence |
AB920580 |
1312 |
|
1571716 tax ID tax ID |
* |
| [Ref.: #20218] |
Streptomyces sp. NTM2 gene for 16S ribosomal RNA, partial sequence |
AB920583 |
1297 |
|
1571719 tax ID tax ID |
* |
| [Ref.: #20218] |
Streptomyces sp. MTMn4 gene for 16S ribosomal RNA, partial sequence |
AB920585 |
1279 |
|
1571721 tax ID tax ID |
* |
| [Ref.: #20218] |
Streptomyces sp. NTRH1 gene for 16S ribosomal RNA, partial sequence |
AB920586 |
1287 |
|
1571722 tax ID tax ID |
* |
| [Ref.: #20218] |
Streptomyces violaceochromogenes strain IFO 13100 16S ribosomal RNA gene, partial sequence |
AY999867 |
1425 |
|
67377 tax ID tax ID |
* |
| [Ref.: #20218] |
Streptomyces violaceochromogenes gene for 16S ribosomal RNA, partial sequence, strain: JCM 4530 |
D44215 |
121 |
|
67377 tax ID tax ID |
* |
| [Ref.: #20218] |
Streptomyces violaceochromogenes gene for 16S rRNA, partial sequence, strain: NBRC 13100 |
AB184312 |
1477 |
|
67377 tax ID tax ID |
* |
|
|
Genome sequence information: |
|
|
Sequence accession description |
Seq. accession number |
Assembly level |
Sequence database |
Associated NCBI tax ID |
|
| [Ref.: #66792] |
Streptomyces violaceochromogenes JCM 4530 |
GCA_014650235 |
scaffold |
|
67377 tax ID tax ID |
* |
| [Ref.: #66792] |
Streptomyces violaceochromogenes strain JCM 4530 |
67377.3 |
wgs |
|
67377 tax ID tax ID |
* |
|
|
-
Availability in culture collections
 External links External links
| [Ref.: #9245] |
Culture collection no. |
DSM 40181, ATCC 19932, ATCC 25512, CBS 654.69, IFO 13100, INA 425, ISP 5181, NBRC 13100, RIA 1292, JCM 4530, CGMCC 4.1753, KCTC 19974, NRRL B-5427, VKM Ac-581 |
| [Ref.: #85147] |
 SI-ID 45813 SI-ID 45813
|
* |
 |
Literature: |
|
Topic |
Title |
Authors |
Journal |
DOI |
Year |
|
| Pathogenicity |
Nybomycin-producing Streptomyces isolated from carpenter ant Camponotus vagus. |
Zakalyukina YV, Birykov MV, Lukianov DA, Shiriaev DI, Komarova ES, Skvortsov DA, Kostyukevich Y, Tashlitsky VN, Polshakov VI, Nikolaev E, Sergiev PV, Osterman IA |
Biochimie |
10.1016/j.biochi.2019.02.010 |
2019 |
* |
|
 References References-
| #9245 |
Leibniz Institut DSMZ-Deutsche Sammlung von Mikroorganismen und Zellkulturen GmbH ; Curators of the DSMZ;
DSM 40181
|
-
-
-
-
-
| #67770 |
Japan Collection of Microorganism (JCM) ; Curators of the JCM;
|
-
| #68368 |
Automatically annotated from API 20E .
|
-
-
-
| #85147 |
Reimer, L.C., Lissin, A.,Schober, I., Witte,J.F., Podstawka, A., Lüken, H., Bunk, B.,Overmann, J.:
StrainInfo: A central database for resolving microbial strain identifiers
.
(
DOI 10.60712/SI-ID45813.1 )
|
- * These data were automatically processed and therefore are not curated
Change proposal
Successfully sent
|
|
Name and taxonomic classification

Morphology

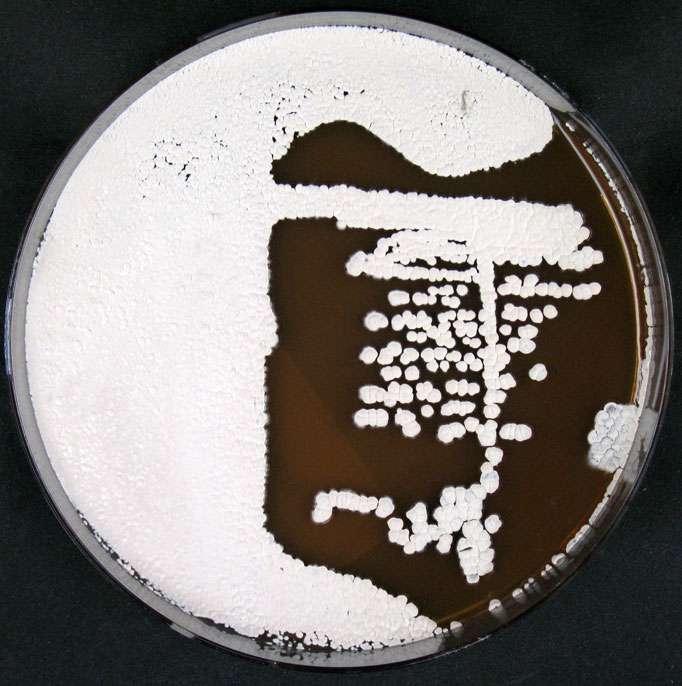
Culture and growth conditions

Physiology and metabolism

Isolation, sampling and environmental information

Safety information

Sequence information

Genome-based predictions

External links

References






