| [Ref.: #18224] |
Culture collection no. |
DSM 26447, BCRC 80167, KCTC 23306 |
| [Ref.: #86917] |
 SI-ID 402256 SI-ID 402256
|
* |
 |
Literature: |
|
Topic |
Title |
Authors |
Journal |
DOI |
Year |
|
| Phylogeny |
Aquabacterium pictum sp. nov., the first aerobic bacteriochlorophyll a-containing fresh water bacterium in the genus Aquabacterium of the class Betaproteobacteria. |
Hirose S, Tank M, Hara E, Tamaki H, Mori K, Takaichi S, Haruta S, Hanada S |
Int J Syst Evol Microbiol |
10.1099/ijsem.0.003798 |
2020 |
* |
| Phylogeny |
Aquabacterium olei sp. nov., an oil-degrading bacterium isolated from oil-contaminated soil. |
Pham VHT, Jeong SW, Kim J |
Int J Syst Evol Microbiol |
10.1099/ijsem.0.000458 |
2015 |
* |
| Phylogeny |
Aquabacterium limnoticum sp. nov., isolated from a freshwater spring. |
Chen WM, Cho NT, Yang SH, Arun AB, Young CC, Sheu SY |
Int J Syst Evol Microbiol |
10.1099/ijs.0.030635-0 |
2011 |
* |
|
 References References-
| #18224 |
Leibniz Institut DSMZ-Deutsche Sammlung von Mikroorganismen und Zellkulturen GmbH ; Curators of the DSMZ;
DSM 26447
|
-
-
-
| #67771 |
Korean Collection for Type Cultures (KCTC) ; Curators of the KCTC;
|
-
| #86917 |
Reimer, L.C., Lissin, A.,Schober, I., Witte,J.F., Podstawka, A., Lüken, H., Bunk, B.,Overmann, J.:
StrainInfo: A central database for resolving microbial strain identifiers
.
(
DOI 10.60712/SI-ID402256.1 )
|
-
- * These data were automatically processed and therefore are not curated
Change proposal
Successfully sent
|
Name and taxonomic classification

Morphology

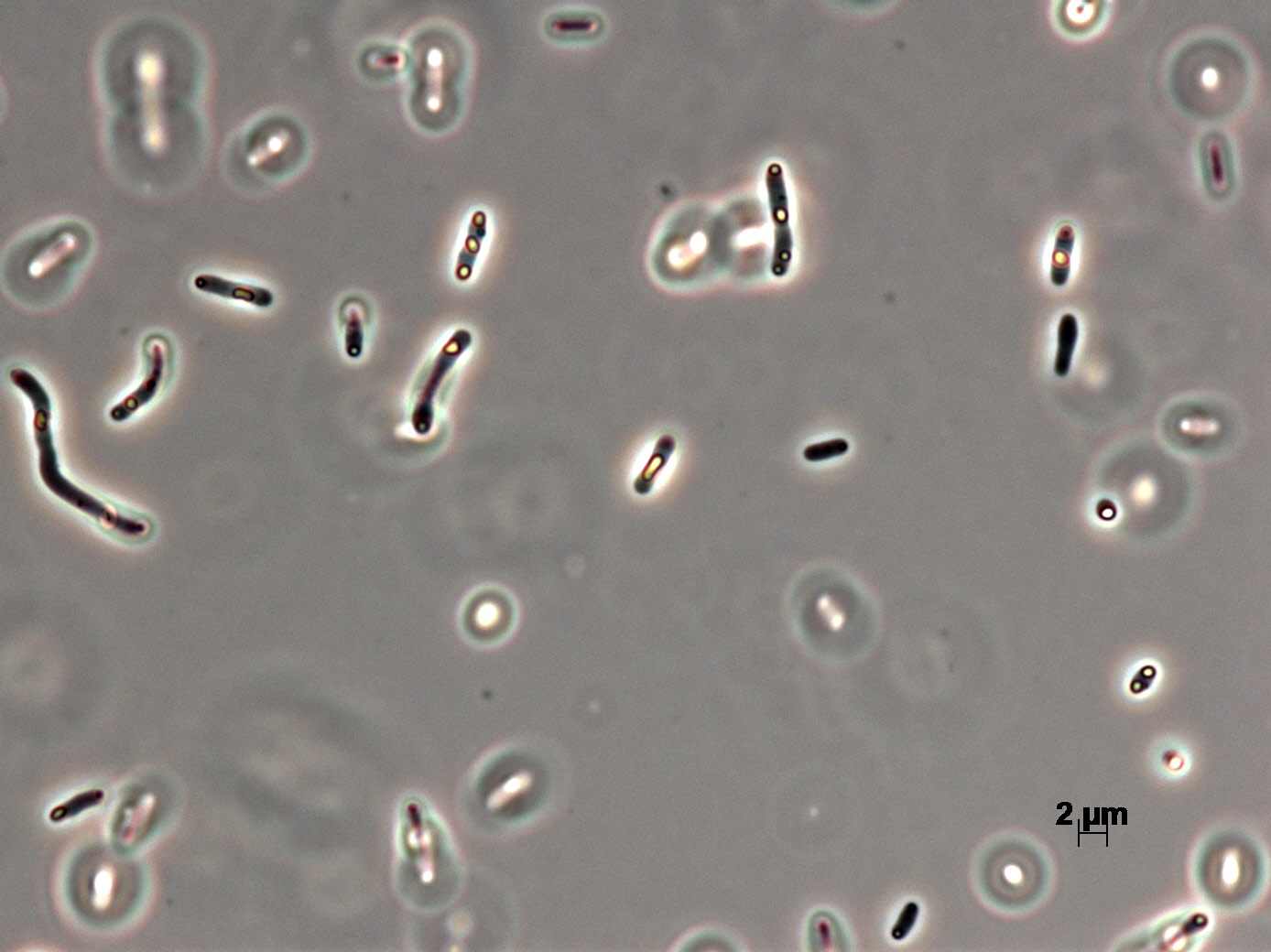
Culture and growth conditions

Physiology and metabolism

Isolation, sampling and environmental information

Safety information

Sequence information

External links

References






