Frankia canadensis ARgP5AG is a bacterium that was isolated from root nodule on an Alnus incana subsp. rugosa shrub.
- 16S sequence
- Bacteria
-
Information on the name and the taxonomic classification.
Name and taxonomic classification

|
-
Information on morphological and physiological properties
Morphology

-

[Ref.: #66793] Caption: electron microscopic image [Ref.: #66793] Intellectual property rights: © HZI/Manfred Rohde -
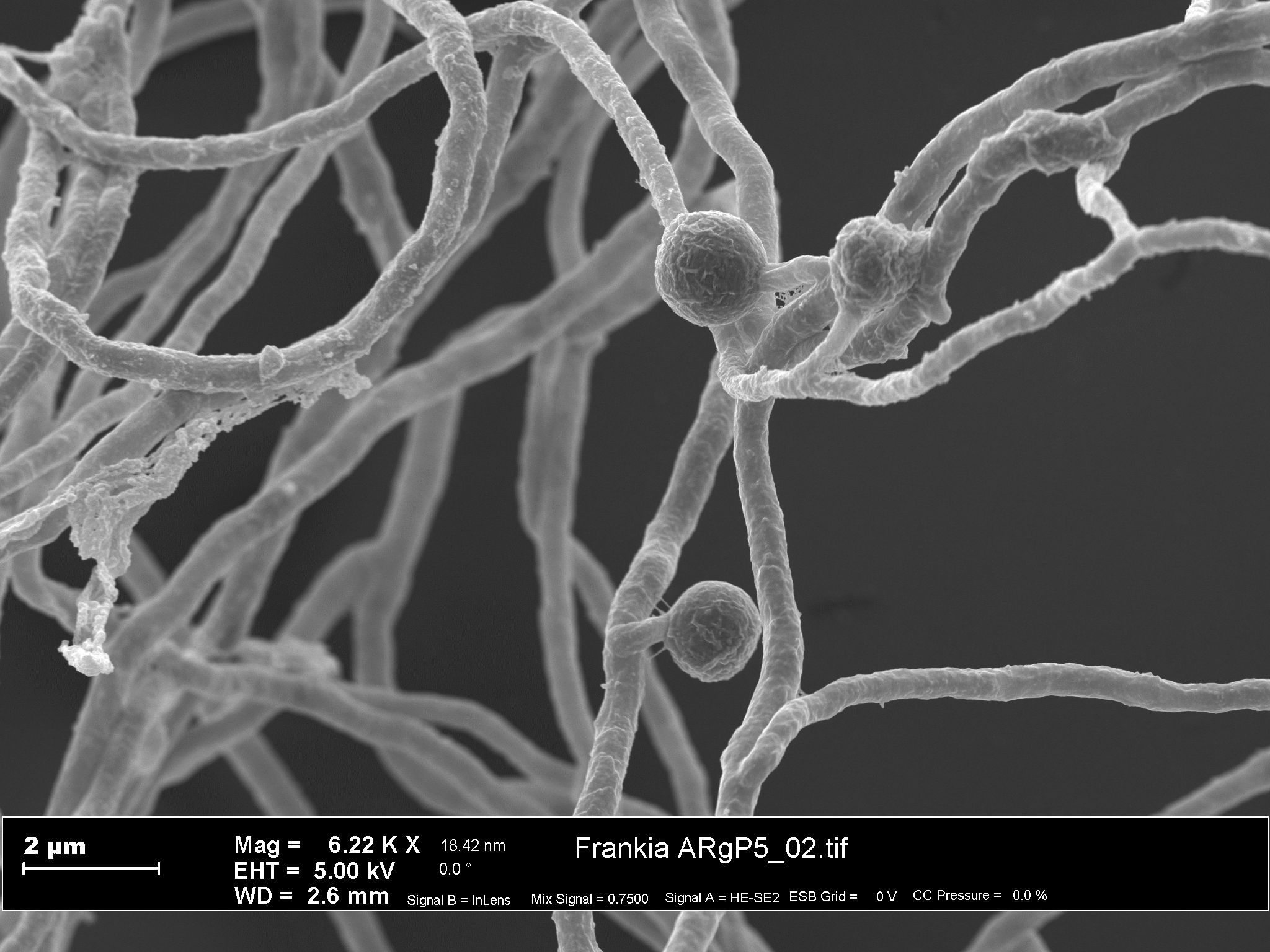
[Ref.: #66793] Caption: electron microscopic image [Ref.: #66793] Intellectual property rights: © HZI/Manfred Rohde -

[Ref.: #66793] Caption: electron microscopic image [Ref.: #66793] Intellectual property rights: © HZI/Manfred Rohde
|
-
Information on culture and growth conditions
Culture and growth conditions

| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
-
Information on physiology and metabolism
Physiology and metabolism

| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
-
Information on isolation source, the sampling and environmental conditions
Isolation, sampling and environmental information

-
Information on genomic background e.g. entries in nucleic sequence databass
Sequence information

-
Availability in culture collections
External links

References
-
#20215 Parte, A.C., Sardà Carbasse, J., Meier-Kolthoff, J.P., Reimer, L.C. and Göker, M.: List of Prokaryotic names with Standing in Nomenclature (LPSN) moves to the DSMZ. IJSEM ( DOI 10.1099/ijsem.0.004332 ) -
#65047 Leibniz Institut DSMZ-Deutsche Sammlung von Mikroorganismen und Zellkulturen GmbH ; Curators of the DSMZ; DSM 45898 -
#66668 Philippe Normand, Imen Nouioui, Petar Pujic, Pascale Fournier, Audrey Dubost, Guillaume Schwob, Hans-Peter Klenk, Agnes Nguyen, Danis Abrouk, Aude Herrera-Belaroussi, Joel F. Pothier, Valentin Pflüger, Maria P. Fernandez: Frankia canadensis sp. nov., isolated from root nodules of Alnus incana subspecies rugosa. IJSEM 68: 3001 - 3011 2018 ( DOI 10.1099/ijsem.0.002939 , PubMed 30059001 ) -
#66793 Mukherjee et al.: GEBA: 1,003 reference genomes of bacterial and archaeal isolates expand coverage of the tree of life. 35: 676 - 683 2017 ( DOI 10.1038/nbt.3886 , PubMed 28604660 ) -
#66794 Antje Chang, Lisa Jeske, Sandra Ulbrich, Julia Hofmann, Julia Koblitz, Ida Schomburg, Meina Neumann-Schaal, Dieter Jahn, Dietmar Schomburg: BRENDA, the ELIXIR core data resource in 2021: new developments and updates. Nucleic Acids Res. 49: D498 - D508 2020 ( DOI 10.1093/nar/gkaa1025 , PubMed 33211880 ) -
#111176 Reimer, L.C., Lissin, A.,Schober, I., Witte,J.F., Podstawka, A., Lüken, H., Bunk, B.,Overmann, J.: StrainInfo: A central database for resolving microbial strain identifiers . ( DOI 10.60712/SI-ID400479.1 ) - * These data were automatically processed and therefore are not curated
| |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||








