| Metabolism |
Chemical diversity of labdane-type bicyclic diterpene biosynthesis in Actinomycetales microorganisms. |
Yamada Y, Komatsu M, Ikeda H |
J Antibiot (Tokyo) |
10.1038/ja.2015.147 |
2016 |
* |
| Metabolism |
Screening of wild type Streptomyces isolates able to overproduce clavulanic acid. |
Viana Marques DA, Santos-Ebinuma Vde C, de Oliveira PM, Lima GM, Araujo JM, Lima-Filho JL, Converti A, Pessoa-Junior A, Porto AL |
Braz J Microbiol |
10.1590/s1517-83822014000300022 |
2014 |
* |
| Phylogeny |
Beta-lactam antibiotic biosynthetic genes have been conserved in clusters in prokaryotes and eukaryotes. |
Smith DJ, Burnham MK, Bull JH, Hodgson JE, Ward JM, Browne P, Brown J, Barton B, Earl AJ, Turner G |
EMBO J |
10.1002/j.1460-2075.1990.tb08168.x |
1990 |
* |
| Metabolism |
Clavulanic acid: a beta-lactamase-inhiting beta-lactam from Streptomyces clavuligerus. |
Reading C, Cole M |
Antimicrob Agents Chemother |
10.1128/AAC.11.5.852 |
1977 |
* |
| Biotechnology |
Environmental Factors Modulate the Role of orf21 Sigma Factor in Clavulanic Acid Production in Streptomyces Clavuligerus ATCC27064. |
Patino LF, Aguirre-Hoyos V, Pinilla LI, Toro LF, Rios-Estepa R |
Bioengineering (Basel) |
10.3390/bioengineering9020078 |
2022 |
* |
| Genetics |
Teleocidin-producing genotype of Streptomyces clavuligerus ATCC 27064. |
Pivk Lukancic P, Drcar T, Bruccoleri R, Crnugelj M, Mrak P |
Appl Microbiol Biotechnol |
10.1007/s00253-022-11805-5 |
2022 |
* |
| Metabolism |
Proteomic analysis of a hom-disrupted, cephamycin C overproducing Streptomyces clavuligerus. |
Unsaldi E, Kurt-Kizildogan A, Ozcan S, Becher D, Voigt B, Aktas C, Ozcengiz G |
Protein Pept Lett |
10.2174/0929866527666200723163655 |
2021 |
* |
| Transcriptome |
Comparative Transcriptome Analysis of Streptomyces Clavuligerus in Response to Favorable and Restrictive Nutritional Conditions. |
Pinilla L, Toro LF, Laing E, Alzate JF, Rios-Estepa R |
Antibiotics (Basel) |
10.3390/antibiotics8030096 |
2019 |
* |
| Metabolism |
The CagRS Two-Component System Regulates Clavulanic Acid Metabolism via Multiple Pathways in Streptomyces clavuligerus F613-1. |
Fu J, Qin R, Zong G, Liu C, Kang N, Zhong C, Cao G |
Front Microbiol |
10.3389/fmicb.2019.00244 |
2019 |
* |
| Metabolism |
Transcriptional Studies on a Streptomyces clavuligerus oppA2 Deletion Mutant: N-Acetylglycyl-Clavaminic Acid Is an Intermediate of Clavulanic Acid Biosynthesis. |
Alvarez-Alvarez R, Rodriguez-Garcia A, Martinez-Burgo Y, Martin JF, Liras P |
Appl Environ Microbiol |
10.1128/AEM.01701-18 |
2018 |
* |
| Genetics |
Genome-wide identification and characterization of reference genes with different transcript abundances for Streptomyces coelicolor. |
Li S, Wang W, Li X, Fan K, Yang K |
Sci Rep |
10.1038/srep15840 |
2015 |
* |
| Metabolism |
The Pathway-Specific Regulator ClaR of Streptomyces clavuligerus Has a Global Effect on the Expression of Genes for Secondary Metabolism and Differentiation. |
Martinez-Burgo Y, Alvarez-Alvarez R, Rodriguez-Garcia A, Liras P |
Appl Environ Microbiol |
10.1128/AEM.00916-15 |
2015 |
* |
| Metabolism |
Heterologous expression of Streptomyces clavuligerus ATCC 27064 cephamycin C gene cluster. |
Martinez-Burgo Y, Alvarez-Alvarez R, Perez-Redondo R, Liras P |
J Biotechnol |
10.1016/j.jbiotec.2014.06.002 |
2014 |
* |
| Metabolism |
Clavulanic acid production by the MMS 150 mutant obtained from wild type Streptomyces clavuligerus ATCC 27064. |
da Silva Vasconcelos E, de Lima VA, Goto LS, Cruz-Hernandez IL, Hokka CO |
Braz J Microbiol |
10.1590/S1517-83822014005000005 |
2014 |
* |
| Metabolism |
Transcriptomic analysis of Streptomyces clavuligerus DeltaccaR::tsr: effects of the cephamycin C-clavulanic acid cluster regulator CcaR on global regulation. |
Alvarez-Alvarez R, Rodriguez-Garcia A, Santamarta I, Perez-Redondo R, Prieto-Dominguez A, Martinez-Burgo Y, Liras P |
Microb Biotechnol |
10.1111/1751-7915.12109 |
2014 |
* |
| Metabolism |
Holomycin, a dithiolopyrrolone compound produced by Streptomyces clavuligerus. |
Liras P |
Appl Microbiol Biotechnol |
10.1007/s00253-013-5410-z |
2013 |
* |
| Metabolism |
Coordination of glycerol utilization and clavulanic acid biosynthesis to improve clavulanic acid production in Streptomyces clavuligerus. |
Guo D, Zhao Y, Yang K |
Sci China Life Sci |
10.1007/s11427-013-4507-z |
2013 |
* |
| Metabolism |
Transcriptional analysis and proteomics of the holomycin gene cluster in overproducer mutants of Streptomyces clavuligerus. |
Robles-Reglero V, Santamarta I, Alvarez-Alvarez R, Martin JF, Liras P |
J Biotechnol |
10.1016/j.jbiotec.2012.09.017 |
2012 |
* |
| Metabolism |
Prediction of the pho regulon in Streptomyces clavuligerus DSM 738. |
Salehghamari E, Hamedi J, Elahi E, Sepehrizadeh Z, Sadeghi M, Muth G |
New Microbiol |
|
2012 |
* |
| Phylogeny |
Molecular Analysis of the Clavulanic Acid Regulatory Gene Isolated from an Iranian Strain of Streptomyces Clavuligerus , PTCC 1709. |
Hojati Z, Salehi Z, Motovali-Bashi M, Korbekandi H, Jami S |
Cell J |
|
2011 |
* |
| Metabolism |
Characterization of DNA-binding sequences for CcaR in the cephamycin-clavulanic acid supercluster of Streptomyces clavuligerus. |
Santamarta I, Lopez-Garcia MT, Kurt A, Nardiz N, Alvarez-Alvarez R, Perez-Redondo R, Martin JF, Liras P |
Mol Microbiol |
10.1111/j.1365-2958.2011.07743.x |
2011 |
* |
| Metabolism |
A rhodanese-like protein is highly overrepresented in the mutant S. clavuligerus oppA2::aph: effect on holomycin and other secondary metabolites production. |
Nardiz N, Santamarta I, Lorenzana LM, Martin JF, Liras P |
Microb Biotechnol |
10.1111/j.1751-7915.2010.00222.x |
2010 |
* |
| Metabolism |
Homologous expression of aspartokinase (ask) gene in Streptomyces clavuligerus and its hom-deleted mutant: effects on cephamycin C production. |
Ozcengiz G, Okay S, Unsaldi E, Taskin B, Liras P, Piret J |
Bioeng Bugs |
10.4161/bbug.1.3.11244 |
2010 |
* |
| Enzymology |
Characterization of the tunicamycin gene cluster unveiling unique steps involved in its biosynthesis. |
Chen W, Qu D, Zhai L, Tao M, Wang Y, Lin S, Price NP, Deng Z |
Protein Cell |
10.1007/s13238-010-0127-6 |
2010 |
* |
| Metabolism |
Identification of the gene cluster for the dithiolopyrrolone antibiotic holomycin in Streptomyces clavuligerus. |
Li B, Walsh CT |
Proc Natl Acad Sci U S A |
10.1073/pnas.1014140107 |
2010 |
* |
| Genetics |
Draft genome sequence of Streptomyces clavuligerus NRRL 3585, a producer of diverse secondary metabolites. |
Song JY, Jeong H, Yu DS, Fischbach MA, Park HS, Kim JJ, Seo JS, Jensen SE, Oh TK, Lee KJ, Kim JF |
J Bacteriol |
10.1128/JB.00859-10 |
2010 |
* |
| Enzymology |
Superoxide dismutase (SOD) genes in Streptomyces peucetius: effects of SODs on secondary metabolites production. |
Kanth BK, Jnawali HN, Niraula NP, Sohng JK |
Microbiol Res |
10.1016/j.micres.2010.07.003 |
2010 |
* |
| Metabolism |
Role of sigma-factor (orf21) in clavulanic acid production in Streptomyces clavuligerus NRRL3585. |
Jnawali HN, Liou K, Sohng JK |
Microbiol Res |
10.1016/j.micres.2010.07.005 |
2010 |
* |
| Genetics |
The sequence of a 1.8-mb bacterial linear plasmid reveals a rich evolutionary reservoir of secondary metabolic pathways. |
Medema MH, Trefzer A, Kovalchuk A, van den Berg M, Muller U, Heijne W, Wu L, Alam MT, Ronning CM, Nierman WC, Bovenberg RA, Breitling R, Takano E |
Genome Biol Evol |
10.1093/gbe/evq013 |
2010 |
* |
| Biotechnology |
Overproduction of Clavulanic Acid by UV Mutagenesis of Streptomyces clavuligerus. |
Korbekandi H, Darkhal P, Hojati Z, Abedi D, Hamedi J, Pourhosein M |
Iran J Pharm Res |
|
2010 |
* |
| Metabolism |
Enhancement of clavulanic acid production by expressing regulatory genes in gap gene deletion mutant of Streptomyces clavuligerus NRRL3585. |
Jnawali HN, Lee HC, Sohng JK |
J Microbiol Biotechnol |
JMB020-01-20 |
2010 |
* |
| Metabolism |
[Evaluation of the activities of two promoters in Streptomycetes by reporter gene method]. |
Li J, Xiang S, Yang X, Yang K |
Wei Sheng Wu Xue Bao |
|
2009 |
* |
| Metabolism |
The enigmatic lack of glucose utilization in Streptomyces clavuligerus is due to inefficient expression of the glucose permease gene. |
Perez-Redondo R, Santamarta I, Bovenberg R, Martin JF, Liras P |
Microbiology (Reading) |
10.1099/mic.0.035840-0 |
2010 |
* |
| Metabolism |
Chapter 16. Enzymology of beta-lactam compounds with cephem structure produced by actinomycete. |
Liras P, Demain AL |
Methods Enzymol |
10.1016/S0076-6879(09)04816-2 |
2009 |
* |
| Metabolism |
Glycerol utilization gene cluster in Streptomyces clavuligerus. |
Banos S, Perez-Redondo R, Koekman B, Liras P |
Appl Environ Microbiol |
10.1128/AEM.00181-09 |
2009 |
* |
| Genetics |
A two-component regulatory system involved in clavulanic acid production. |
Jnawali HN, Oh TJ, Liou K, Park BC, Sohng JK |
J Antibiot (Tokyo) |
10.1038/ja.2008.92 |
2008 |
* |
| Metabolism |
An approach to strain improvement and enhanced production of clavulanic acid in Streptomyces clavuligerus. |
Kim SJ, Kim JO, Shin CH, Park HW, Kim CW |
Biosci Biotechnol Biochem |
10.1271/bbb.80569 |
2009 |
* |
| Biotechnology |
A gene located downstream of the clavulanic acid gene cluster in Streptomyces clavuligerus ATCC 27064 encodes a putative response regulator that affects clavulanic acid production. |
Song JY, Kim ES, Kim DW, Jensen SE, Lee KJ |
J Ind Microbiol Biotechnol |
10.1007/s10295-008-0499-2 |
2008 |
* |
| Metabolism |
Functional effects of increased copy number of the gene encoding proclavaminate amidino hydrolase on clavulanic acid production in Streptomyces clavuligerus ATCC 27064. |
Song JY, Kim ES, Kim DW, Jensen SE, Lee KJ |
J Microbiol Biotechnol |
7169 |
2008 |
* |
| Metabolism |
Enhancement of clavulanic acid by replicative and integrative expression of ccaR and cas2 in Streptomyces clavuligerus NRRL3585. |
Hung TV, Malla S, Park BC, Liou K, Lee HC, Sohng JK |
J Microbiol Biotechnol |
|
2007 |
* |
| Enzymology |
Targeted disruption of homoserine dehydrogenase gene and its effect on cephamycin C production in Streptomyces clavuligerus. |
Yilmaz EI, Caydasi AK, Ozcengiz G |
J Ind Microbiol Biotechnol |
10.1007/s10295-007-0259-8 |
2007 |
* |
| Metabolism |
Connecting primary and secondary metabolism: AreB, an IclR-like protein, binds the ARE(ccaR) sequence of S. clavuligerus and modulates leucine biosynthesis and cephamycin C and clavulanic acid production. |
Santamarta I, Lopez-Garcia MT, Perez-Redondo R, Koekman B, Martin JF, Liras P |
Mol Microbiol |
10.1111/j.1365-2958.2007.05937.x |
2007 |
* |
| Metabolism |
A statistical approach using L(25) orthogonal array method to study fermentative production of clavulanic acid by Streptomyces clavuligerus MTCC 1142. |
Saudagar PS, Singhal RS |
Appl Biochem Biotechnol |
10.1007/s12010-007-9030-x |
2007 |
* |
| Genetics |
Prediction and functional analysis of the replication origin of the linear plasmid pSCL2 in Streptomyces clavuligerus. |
Wu W, Leblanc SK, Piktel J, Jensen SE, Roy KL |
Can J Microbiol |
10.1139/w05-126 |
2006 |
* |
| Metabolism |
Cephamycin C production is regulated by relA and rsh genes in Streptomyces clavuligerus ATCC27064. |
Jin W, Kim HK, Kim JY, Kang SG, Lee SH, Lee KJ |
J Biotechnol |
10.1016/j.jbiotec.2004.06.010 |
2004 |
* |
| Enzymology |
Cloning, characterization and heterologous expression of the aspartokinase and aspartate semialdehyde dehydrogenase genes of cephamycin C-producer Streptomyces clavuligerus. |
Tunca S, Yilmaz EI, Piret J, Liras P, Ozcengiz G |
Res Microbiol |
10.1016/j.resmic.2004.03.007 |
2004 |
* |
| Metabolism |
CcaR is an autoregulatory protein that binds to the ccaR and cefD-cmcI promoters of the cephamycin C-clavulanic acid cluster in Streptomyces clavuligerus. |
Santamarta I, Rodriguez-Garcia A, Perez-Redondo R, Martin JF, Liras P |
J Bacteriol |
10.1128/JB.184.11.3106-3113.2002 |
2002 |
* |
| Enzymology |
Cloning, heterologous expression and purification of an isocitrate lyase from Streptomyces clavuligerus NRRL 3585. |
Soh BS, Loke P, Sim TS |
Biochim Biophys Acta |
10.1016/s0167-4781(01)00309-8 |
2001 |
* |
| Enzymology |
Malate synthase from Streptomyces clavuligerus NRRL3585: cloning, molecular characterization and its control by acetate. |
Chan M, Sim TS |
Microbiology (Reading) |
10.1099/00221287-144-11-3229 |
1998 |
* |
| Metabolism |
Lysine epsilon-aminotransferase, the initial enzyme of cephalosporin biosynthesis in actinomycetes. |
Rius N, Demain AL |
J Microbiol Biotechnol |
|
1997 |
* |
| Metabolism |
Effect of water activity on production of beta-lactam antibiotics by Streptomyces clavuligerus in submerged culture. |
Cochet N, Demain AL |
J Appl Bacteriol |
10.1111/j.1365-2672.1996.tb03228.x |
1996 |
* |
| Genetics |
Giant linear plasmids of beta-lactam antibiotic producing Streptomyces. |
Netolitzky DJ, Wu X, Jensen SE, Roy KL |
FEMS Microbiol Lett |
10.1016/0378-1097(95)00230-3 |
1995 |
* |
| Enzymology |
Cloning of a Streptomyces clavuligerus DNA fragment encoding the cephalosporin 7 alpha-hydroxylase and its expression in Streptomyces lividans. |
Xiao X, Hintermann G, Hausler A, Barker PJ, Foor F, Demain AL, Piret J |
Antimicrob Agents Chemother |
10.1128/AAC.37.1.84 |
1993 |
* |
| Metabolism |
Characterization of an inducible transport system for glycerol in Streptomyces clavuligerus. Repression by L-serine. |
Minambres B, Reglero A, Luengo JM |
J Antibiot (Tokyo) |
10.7164/antibiotics.45.269 |
1992 |
* |
| Enzymology |
Purification and characterization of clavaminate synthase from Streptomyces clavuligerus: an unusual oxidative enzyme in natural product biosynthesis. |
Salowe SP, Marsh EN, Townsend CA |
Biochemistry |
10.1021/bi00479a023 |
1990 |
* |
| Enzymology |
Characterization and complementation of a cephalosporin-deficient mutant of Streptomyces clavuligerus NRRL 3585. |
Piret J, Resendiz B, Mahro B, Zhang JY, Serpe E, Romero J, Connors N, Demain AL |
Appl Microbiol Biotechnol |
10.1007/BF00173728 |
1990 |
* |
| Enzymology |
Regulation of ACV synthetase: biosynthesis and action. |
Zhang JY, Demain AL |
Chin J Biotechnol |
|
1990 |
* |
| Enzymology |
Oxygen derepresses deacetoxycephalosporin C synthase and increases the conversion of penicillin N to cephamycin C in Streptomyces clavuligerus. |
Rollins MJ, Jensen SE, Wolfe S, Westlake DW |
Enzyme Microb Technol |
10.1016/0141-0229(90)90178-s |
1990 |
* |
| Metabolism |
Characterization of sugar uptake in wild-type Streptomyces clavuligerus, which is impaired in glucose uptake, and in a glucose-utilizing mutant. |
Garcia-Dominguez M, Martin JF, Liras P |
J Bacteriol |
10.1128/jb.171.12.6808-6814.1989 |
1989 |
* |
| Metabolism |
Solid state fermentation for cephalosporin production by Streptomyces clavuligerus and Cephalosporium acremonium. |
Jermini MF, Demain AL |
Experientia |
10.1007/BF01950159 |
1989 |
* |
| Enzymology |
Ro 22-5417, a new clavam antibiotic from Streptomyces clavuligerus. I. Discovery and biological activity. |
Pruess DL, Kellett M |
J Antibiot (Tokyo) |
10.7164/antibiotics.36.208 |
1983 |
* |
| Metabolism |
High performance liquid chromatographic assay of cyclization activity in cell-free systems from Streptomyces clavuligerus. |
Jensen SE, Westlake DW, Wolfe S |
J Antibiot (Tokyo) |
10.7164/antibiotics.35.1026 |
1982 |
* |
| Metabolism |
Molecular mechanism of GylR-mediated regulation of glycerol metabolism in Streptomyces clavuligerus NRRL 3585. |
Zhang C, Zhao Y, Li Z, Wang W, Huang Y, Pan G, Fan K |
Front Microbiol |
10.3389/fmicb.2022.1078293 |
2022 |
* |
| Proteome |
An integrative-omics analysis of an industrial clavulanic acid-overproducing Streptomyces clavuligerus. |
Kurt-Kizildogan A, Celik G, Unsaldi E, Ozcan S, Ayaz-Guner S, Ozcengiz G |
Appl Microbiol Biotechnol |
10.1007/s00253-022-12098-4 |
2022 |
* |
|
Genome editing reveals that pSCL4 is required for chromosome linearity in Streptomyces clavuligerus. |
Gomez-Escribano JP, Algora Gallardo L, Bozhuyuk KAJ, Kendrew SG, Huckle BD, Crowhurst NA, Bibb MJ, Collis AJ, Micklefield J, Herron PR, Wilkinson B |
Microb Genom |
10.1099/mgen.0.000669 |
2021 |
* |
|
Elucidating the Regulatory Elements for Transcription Termination and Posttranscriptional Processing in the Streptomyces clavuligerus Genome. |
Hwang S, Lee N, Choe D, Lee Y, Kim W, Jeong Y, Cho S, Palsson BO, Cho BK |
mSystems |
10.1128/mSystems.01013-20 |
2021 |
* |
|
The two-component system CepRS regulates the cephamycin C biosynthesis in Streptomyces clavuligerus F613-1. |
Fu J, Qin R, Zong G, Zhong C, Zhang P, Kang N, Qi X, Cao G |
AMB Express |
10.1186/s13568-019-0844-z |
2019 |
* |
|
Activation of Secondary Metabolite Gene Clusters in Streptomyces clavuligerus by the PimM Regulator of Streptomyces natalensis. |
Martinez-Burgo Y, Santos-Aberturas J, Rodriguez-Garcia A, Barreales EG, Tormo JR, Truman AW, Reyes F, Aparicio JF, Liras P |
Front Microbiol |
10.3389/fmicb.2019.00580 |
2019 |
* |
|
A putative antimicrobial peptide from Hymenoptera in the megaplasmid pSCL4 of Streptomyces clavuligerus ATCC 27064 reveals a singular case of horizontal gene transfer with potential applications. |
Ayala-Ruano S, Santander-Gordon D, Tejera E, Perez-Castillo Y, Armijos-Jaramillo V |
Ecol Evol |
10.1002/ece3.4924 |
2019 |
* |
|
Impacts of horizontal gene transfer on the compact genome of the clavulanic acid-producing Streptomyces strain F613-1. |
Li J, Zhao Z, Zhong W, Zhong C, Zong G, Fu J, Cao G |
3 Biotech |
10.1007/s13205-018-1498-2 |
2018 |
* |
|
A method for investigating the stereochemical course of terpene cyclisations. |
Rabe P, Rinkel J, Klapschinski TA, Barra L, Dickschat JS |
Org Biomol Chem |
10.1039/c5ob01998b |
2016 |
* |
Name and taxonomic classification

Morphology

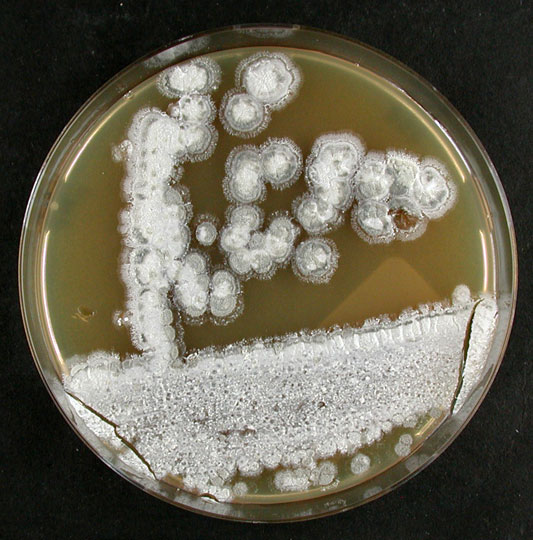

Culture and growth conditions

Physiology and metabolism

Isolation, sampling and environmental information

Safety information

Sequence information

Genome-based predictions

External links

References








