| [Ref.: #18181] |
Sample type/isolated from |
roots of diseased white mulberry plant Morus alba L. |
| [Ref.: #18181] |
Host species |
Morus alba |
| [Ref.: #18181] |
Geographic location (country and/or sea, region) |
Zhejiang province, Hangzhou, Tonglu |
| [Ref.: #18181] |
Country |
China |
| [Ref.: #18181] |
Country ISO 3 Code |
CHN |
| [Ref.: #18181] |
Continent |
Asia |
|
* marker position based on {}
|
|
Isolation sources categories |
| #Infection |
- |
- |
| #Host |
#Plants |
#Tree |
| #Host Body-Site |
#Plant |
#Root (Rhizome) |
|
 |
-
Availability in culture collections
 External links External links
| [Ref.: #18181] |
Culture collection no. |
DSM 26271, CGMCC 1.10322, LMG 25706 |
| [Ref.: #73875] |
 SI-ID 369642 SI-ID 369642
|
* |
 |
Literature: |
|
Topic |
Title |
Authors |
Journal |
DOI |
Year |
|
| Phylogeny |
Enterobacter tabaci sp. nov., a novel member of the genus Enterobacter isolated from a tobacco stem. |
Duan YQ, Zhou XK, Di-Yan L, Li QQ, Dang LZ, Zhang YG, Qiu LH, Nimaichand S, Li WJ |
Antonie Van Leeuwenhoek |
10.1007/s10482-015-0569-1 |
2015 |
* |
| Phylogeny |
Enterobacter xiangfangensis sp. nov., isolated from Chinese traditional sourdough, and reclassification of Enterobacter sacchari Zhu et al. 2013 as Kosakonia sacchari comb. nov. |
Gu CT, Li CY, Yang LJ, Huo GC |
Int J Syst Evol Microbiol |
10.1099/ijs.0.064709-0 |
2014 |
* |
| Genetics |
Genome sequence of the Enterobacter mori type strain, LMG 25706, a pathogenic bacterium of Morus alba L. |
Zhu B, Zhang GQ, Lou MM, Tian WX, Li B, Zhou XP, Wang GF, Liu H, Xie GL, Jin GL |
J Bacteriol |
10.1128/JB.05200-11 |
2011 |
* |
| Phylogeny |
Enterobacter mori sp. nov., associated with bacterial wilt on Morus alba L. |
Zhu B, Lou MM, Xie GL, Wang GF, Zhou Q, Wang F, Fang Y, Su T, Li B, Duan YP |
Int J Syst Evol Microbiol |
10.1099/ijs.0.028613-0 |
2011 |
* |
|
First Report of Bacterial Wilt Caused by Enterobacter mori of Strawberry in Beijing, China. |
Ji S, Li H, Zhou Y, Li X, Yan J, Zhang W |
Plant Dis |
10.1094/PDIS-08-22-1895-PDN |
2022 |
* |
|
 References References-
| #18181 |
Leibniz Institut DSMZ-Deutsche Sammlung von Mikroorganismen und Zellkulturen GmbH ; Curators of the DSMZ;
DSM 26271
|
-
-
-
-
| #68368 |
Automatically annotated from API 20E .
|
-
| #69479 |
João F Matias Rodrigues, Janko Tackmann,Gregor Rot, Thomas SB Schmidt, Lukas Malfertheiner, Mihai Danaila,Marija Dmitrijeva, Daniela Gaio, Nicolas Näpflin and Christian von Mering. University of Zurich.:
MicrobeAtlas 1.0 beta
.
|
-
-
-
| #73875 |
Reimer, L.C., Lissin, A.,Schober, I., Witte,J.F., Podstawka, A., Lüken, H., Bunk, B.,Overmann, J.:
StrainInfo: A central database for resolving microbial strain identifiers
.
(
DOI 10.60712/SI-ID369642.1 )
|
-
| #124043 |
Dr. Isabel Schober, Dr. Julia Koblitz:
Data extracted from sequence databases, automatically matched based on designation and taxonomy .
|
-
- * These data were automatically processed and therefore are not curated
Change proposal
Successfully sent
|
|
Name and taxonomic classification

Morphology

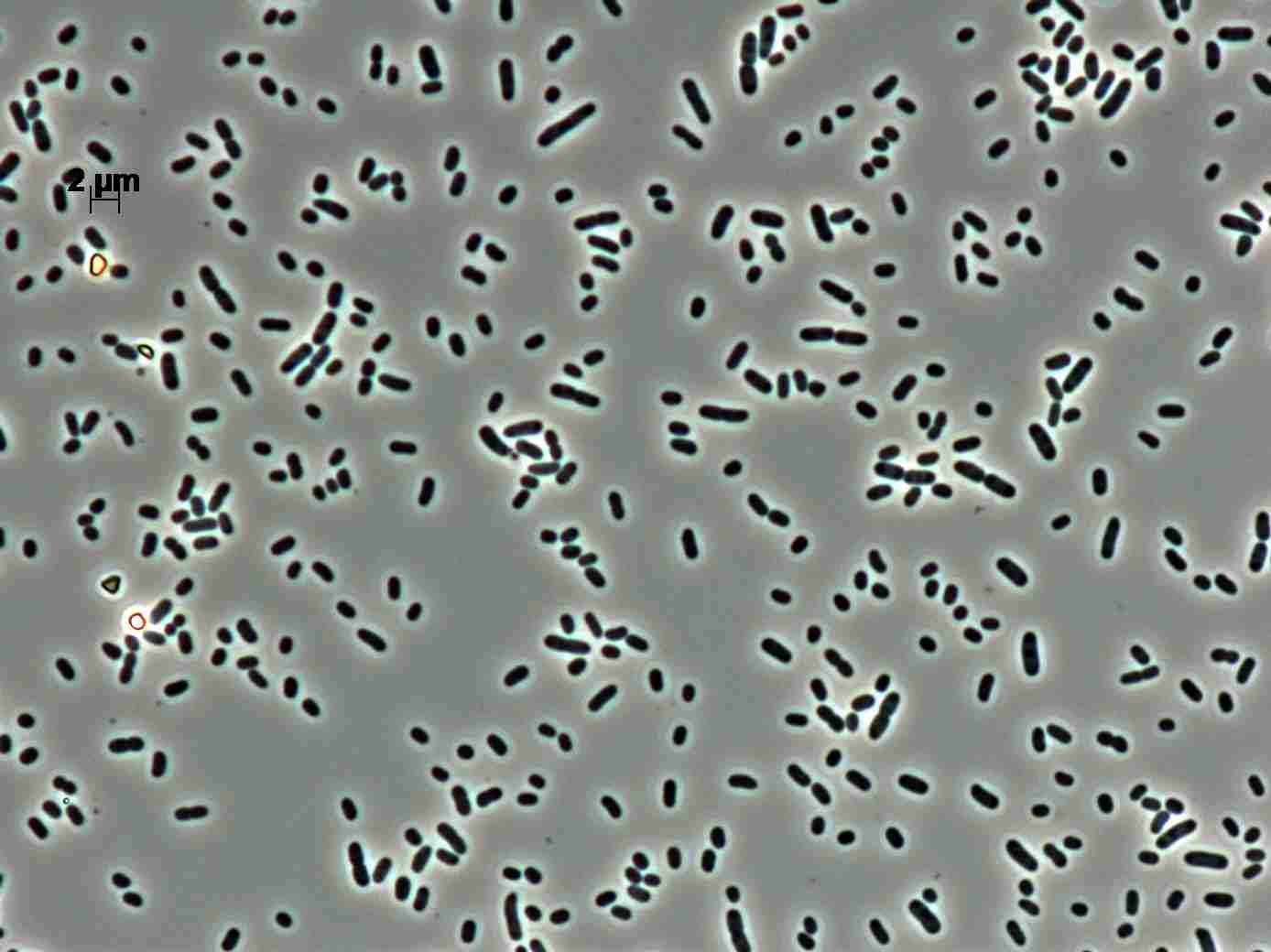
Culture and growth conditions

Physiology and metabolism

Isolation, sampling and environmental information

Safety information

Sequence information

Genome-based predictions

External links

References