| [Ref.: #2888] |
Sample type/isolated from |
human skin |
| |
| [Ref.: #52916] |
Sample type/isolated from |
Human cutaneous |
| [Ref.: #52916] |
Country |
United Kingdom |
| [Ref.: #52916] |
Country ISO 3 Code |
GBR |
| [Ref.: #52916] |
Continent |
Europe |
| |
| [Ref.: #67770] |
Sample type/isolated from |
Human skin |
| |
| [Ref.: #116349] |
Sample type/isolated from |
Human, Skin |
|
* marker position based on {}
|
|
Isolation sources categories |
| #Host |
#Human |
- |
| #Host Body-Site |
#Organ |
#Skin, Nail, Hair |
|
 |
-
Availability in culture collections
 External links External links
| [Ref.: #2888] |
Culture collection no. |
DSM 6922, ATCC 49368, NCFB 2768, CCUG 35685, LMG 19043, JCM 7447, CCM 4123, CECT 4163, CGMCC 1.1939, CIP 103452, IFO 15207, KCTC 3432, NBRC 15207, NCIMB 13130 |
| [Ref.: #72651] |
 SI-ID 12982 SI-ID 12982
|
* |
 |
Literature: |
|
Topic |
Title |
Authors |
Journal |
DOI |
Year |
|
| Metabolism |
Identification and characterization of the channel-forming protein in the cell wall of Corynebacterium amycolatum. |
Soltan Mohammadi N, Mafakheri S, Abdali N, Barcena-Uribarri I, Tauch A, Benz R |
Biochim Biophys Acta |
10.1016/j.bbamem.2013.06.024 |
2013 |
* |
| Phylogeny |
Corynebacterium freneyi sp. nov., alpha-glucosidase-positive strains related to Corynebacterium xerosis. |
Renaud FN, Aubel D, Riegel P, Meugnier H, Bollet C |
Int J Syst Evol Microbiol |
10.1099/00207713-51-5-1723 |
2001 |
* |
| Enzymology |
Comparative chemotaxonomic studies of mycolic acid-free coryneform bacteria of human origin. |
Barreau C, Bimet F, Kiredjian M, Rouillon N, Bizet C |
J Clin Microbiol |
10.1128/jcm.31.8.2085-2090.1993 |
1993 |
* |
|
 References References-
| #2888 |
Leibniz Institut DSMZ-Deutsche Sammlung von Mikroorganismen und Zellkulturen GmbH ; Curators of the DSMZ;
DSM 6922
|
-
-
-
-
| #34280 |
; Curators of the CIP;
|
-
| #52916 |
Culture Collection University of Gothenburg (CCUG) ; Curators of the CCUG;
CCUG 35685
|
-
-
| #67770 |
Japan Collection of Microorganism (JCM) ; Curators of the JCM;
|
-
| #68368 |
Automatically annotated from API 20E .
|
-
| #68379 |
Automatically annotated from API Coryne .
|
-
| #68382 |
Automatically annotated from API zym .
|
-
| #69479 |
João F Matias Rodrigues, Janko Tackmann,Gregor Rot, Thomas SB Schmidt, Lukas Malfertheiner, Mihai Danaila,Marija Dmitrijeva, Daniela Gaio, Nicolas Näpflin and Christian von Mering. University of Zurich.:
MicrobeAtlas 1.0 beta
.
|
-
-
| #72651 |
Reimer, L.C., Lissin, A.,Schober, I., Witte,J.F., Podstawka, A., Lüken, H., Bunk, B.,Overmann, J.:
StrainInfo: A central database for resolving microbial strain identifiers
.
(
DOI 10.60712/SI-ID12982.1 )
|
-
| #116349 |
Collection of Institut Pasteur ; Curators of the CIP;
CIP 103452
|
- * These data were automatically processed and therefore are not curated
Change proposal
Successfully sent
|
|
Name and taxonomic classification

Morphology

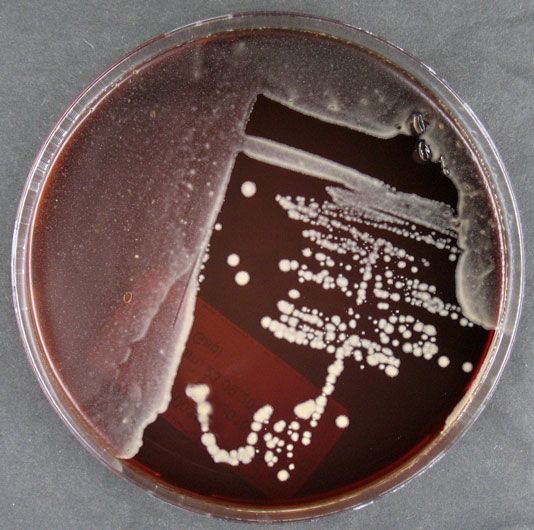
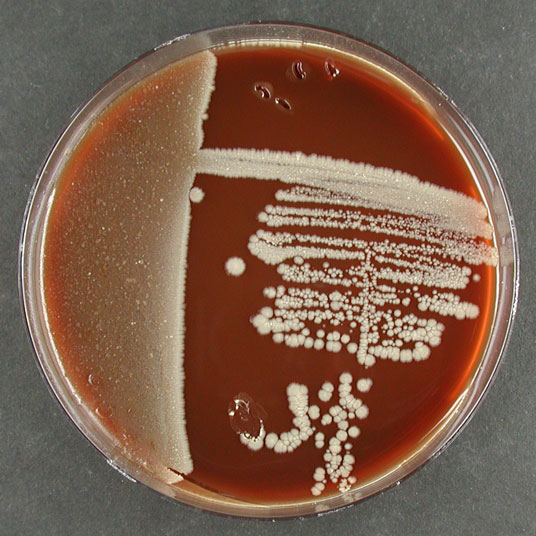
Culture and growth conditions

Physiology and metabolism

Isolation, sampling and environmental information

Safety information

Sequence information

Genome-based predictions

External links

References