| [Ref.: #3206] |
Culture collection no. |
DSM 7534, ATCC 12464, CIP 61.10, NCIMB 547, NCTC 547, CCUG 4855, JCM 8144, BCRC 14552, CCM 5743, CECT 965, CN 3790, NCCB 47070 |
| [Ref.: #72302] |
 SI-ID 13775 SI-ID 13775
|
* |
 |
Literature: |
|
Topic |
Title |
Authors |
Journal |
DOI |
Year |
|
| Phylogeny |
First Comparative Analysis of Clostridium septicum Genomes Provides Insights Into the Taxonomy, Species Genetic Diversity, and Virulence Related to Gas Gangrene. |
Thomas P, Abdel-Glil MY, Subbaiyan A, Busch A, Eichhorn I, Wieler LH, Neubauer H, Pletz M, Seyboldt C |
Front Microbiol |
10.3389/fmicb.2021.771945 |
2021 |
* |
| Phylogeny |
MLST analysis reveals a highly conserved core genome among poultry isolates of Clostridium septicum. |
Neumann AP, Rehberger TG |
Anaerobe |
10.1016/j.anaerobe.2009.01.005 |
2009 |
* |
| Enzymology |
Cloning, nucleotide sequence and expression of a hemolysin gene of Clostridium septicum. |
Imagawa T, Dohi Y, Higashi Y |
FEMS Microbiol Lett |
10.1016/0378-1097(94)90573-8 |
1994 |
* |
|
 References References-
| #3206 |
Leibniz Institut DSMZ-Deutsche Sammlung von Mikroorganismen und Zellkulturen GmbH ; Curators of the DSMZ;
DSM 7534
|
-
-
-
| #37332 |
; Curators of the CIP;
|
-
| #44795 |
Culture Collection University of Gothenburg (CCUG) ; Curators of the CCUG;
CCUG 4855
|
-
-
| #66794 |
Antje Chang, Lisa Jeske, Sandra Ulbrich, Julia Hofmann, Julia Koblitz, Ida Schomburg, Meina Neumann-Schaal, Dieter Jahn, Dietmar Schomburg:
BRENDA, the ELIXIR core data resource in 2021: new developments and updates.
Nucleic Acids Res. 49:
D498 - D508
2020 (
DOI 10.1093/nar/gkaa1025 , PubMed 33211880 )
|
-
| #67770 |
Japan Collection of Microorganism (JCM) ; Curators of the JCM;
|
-
| #68371 |
Automatically annotated from API 50CH acid .
|
-
| #68380 |
Automatically annotated from API rID32A .
|
-
| #69479 |
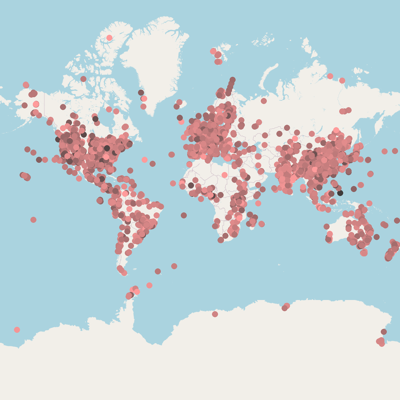
João F Matias Rodrigues, Janko Tackmann,Gregor Rot, Thomas SB Schmidt, Lukas Malfertheiner, Mihai Danaila,Marija Dmitrijeva, Daniela Gaio, Nicolas Näpflin and Christian von Mering. University of Zurich.:
MicrobeAtlas 1.0 beta
.
|
-
-
-
| #72302 |
Reimer, L.C., Lissin, A.,Schober, I., Witte,J.F., Podstawka, A., Lüken, H., Bunk, B.,Overmann, J.:
StrainInfo: A central database for resolving microbial strain identifiers
.
(
DOI 10.60712/SI-ID13775.1 )
|
-
| #122564 |
Collection of Institut Pasteur ; Curators of the CIP;
CIP 61.10
|
- * These data were automatically processed and therefore are not curated
Change proposal
Successfully sent
|
Name and taxonomic classification

Morphology

Culture and growth conditions

Physiology and metabolism

Isolation, sampling and environmental information

Safety information

Sequence information

Genome-based predictions

External links

References