| [Ref.: #20729] |
Culture collection no. |
DSM 26640, CIP 105210, IAM 14812, JCM 21319, ATCC 700781, IFO 16654, NBRC 16654 |
| [Ref.: #88352] |
 SI-ID 44134 SI-ID 44134
|
* |
 |
Literature: |
Only first 10 entries are displayed. Click here to see all.Click here to see only first 10 entries. |
|
Topic |
Title |
Authors |
Journal |
DOI |
Year |
|
| Phylogeny |
Polyphasic taxonomic analysis of Parasedimentitalea marina gen. nov., sp. nov., a psychrotolerant bacterium isolated from deep sea water of the New Britain Trench. |
Ding W, Liu P, Xu Y, Fang J, Cao J |
FEMS Microbiol Lett |
10.1093/femsle/fnaa004 |
2019 |
* |
| Phylogeny |
Phaeobacter porticola sp. nov., an antibiotic-producing bacterium isolated from a sea harbour. |
Breider S, Freese HM, Sproer C, Simon M, Overmann J, Brinkhoff T |
Int J Syst Evol Microbiol |
10.1099/ijsem.0.001879 |
2017 |
* |
| Genetics |
Complete genome sequence of the Phaeobacter gallaeciensis type strain CIP 105210(T) (= DSM 26640(T) = BS107(T)). |
Frank O, Pradella S, Rohde M, Scheuner C, Klenk HP, Goker M, Petersen J |
Stand Genomic Sci |
10.4056/sigs.5179110 |
2014 |
* |
| Phylogeny |
Cribrihabitans neustonicus sp. nov., isolated from coastal surface seawater, and emended description of the genus Cribrihabitans Chen et al. 2014. |
Hameed A, Shahina M, Lin SY, Lai WA, Liu YC, Hsu YH, Young CC |
Int J Syst Evol Microbiol |
10.1099/ijs.0.066142-0 |
2014 |
* |
| Phylogeny |
Molecular and phenotypic analyses reveal the non-identity of the Phaeobacter gallaeciensis type strain deposits CIP 105210T and DSM 17395. |
Buddruhs N, Pradella S, Goker M, Pauker O, Pukall R, Sproer C, Schumann P, Petersen J, Brinkhoff T |
Int J Syst Evol Microbiol |
10.1099/ijs.0.053900-0 |
2013 |
* |
| Phylogeny |
Reclassification of Roseobacter gallaeciensis Ruiz-Ponte et al. 1998 as Phaeobacter gallaeciensis gen. nov., comb. nov., description of Phaeobacter inhibens sp. nov., reclassification of Ruegeria algicola (Lafay et al. 1995) Uchino et al. 1999 as Marinovum algicola gen. nov., comb. nov., and emended descriptions of the genera Roseobacter, Ruegeria and Leisingera. |
Martens T, Heidorn T, Pukall R, Simon M, Tindall BJ, Brinkhoff T |
Int J Syst Evol Microbiol |
10.1099/ijs.0.63724-0 |
2006 |
* |
| Phylogeny |
Paraphaeobacter pallidus gen. nov., sp. nov., isolated from seawater. |
Cai X, Wang Y, Yang X, Liu J, Wu Y, Zhang XH |
Int J Syst Evol Microbiol |
10.1099/ijsem.0.001935 |
2017 |
* |
| Phylogeny |
Seohaeicola nanhaiensis sp. nov., a moderately halophilic bacterium isolated from the benthic sediment of South China Sea. |
Xie BS, Lv XL, Cai M, Tang YQ, Wang YN, Cui HL, Liu XY, Tan Y, Wu XL |
Curr Microbiol |
10.1007/s00284-014-0658-9 |
2014 |
* |
| Phylogeny |
Roseobacter gallaeciensis sp. nov., a new marine bacterium isolated from rearings and collectors of the scallop Pecten maximus. |
Ruiz-Ponte C, Cilia V, Lambert C, Nicolas JL |
Int J Syst Bacteriol |
10.1099/00207713-48-2-537 |
1998 |
* |
|
Adaptation of Surface-Associated Bacteria to the Open Ocean: A Genomically Distinct Subpopulation of Phaeobacter gallaeciensis Colonizes Pacific Mesozooplankton. |
Freese HM, Methner A, Overmann J |
Front Microbiol |
10.3389/fmicb.2017.01659 |
2017 |
* |
|
 References References-
-
| #20729 |
Leibniz Institut DSMZ-Deutsche Sammlung von Mikroorganismen und Zellkulturen GmbH ; Curators of the DSMZ;
DSM 26640
|
-
| #41627 |
; Curators of the CIP;
|
-
-
| #66794 |
Antje Chang, Lisa Jeske, Sandra Ulbrich, Julia Hofmann, Julia Koblitz, Ida Schomburg, Meina Neumann-Schaal, Dieter Jahn, Dietmar Schomburg:
BRENDA, the ELIXIR core data resource in 2021: new developments and updates.
Nucleic Acids Res. 49:
D498 - D508
2020 (
DOI 10.1093/nar/gkaa1025 , PubMed 33211880 )
|
-
| #67770 |
Japan Collection of Microorganism (JCM) ; Curators of the JCM;
|
-
| #68369 |
Automatically annotated from API 20NE .
|
-
| #68382 |
Automatically annotated from API zym .
|
-
| #69479 |
João F Matias Rodrigues, Janko Tackmann,Gregor Rot, Thomas SB Schmidt, Lukas Malfertheiner, Mihai Danaila,Marija Dmitrijeva, Daniela Gaio, Nicolas Näpflin and Christian von Mering. University of Zurich.:
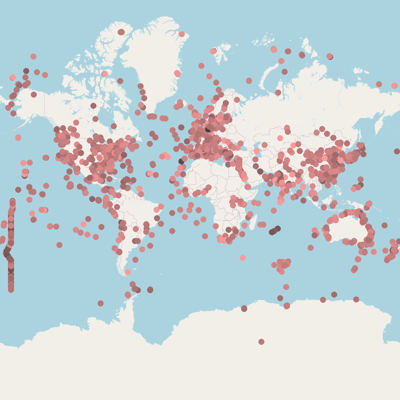
MicrobeAtlas 1.0 beta
.
|
-
-
-
| #88352 |
Reimer, L.C., Lissin, A.,Schober, I., Witte,J.F., Podstawka, A., Lüken, H., Bunk, B.,Overmann, J.:
StrainInfo: A central database for resolving microbial strain identifiers
.
(
DOI 10.60712/SI-ID44134.1 )
|
-
| #119508 |
Collection of Institut Pasteur ; Curators of the CIP;
CIP 105210
|
- * These data were automatically processed and therefore are not curated
Change proposal
Successfully sent
|
Name and taxonomic classification

Morphology

Culture and growth conditions

Physiology and metabolism

Isolation, sampling and environmental information

Safety information

Sequence information

Genome-based predictions

External links

References