| [Ref.: #8672] |
Sample type/isolated from |
human skin |
| |
| [Ref.: #44974] |
Sample type/isolated from |
Human skin |
| [Ref.: #44974] |
Geographic location (country and/or sea, region) |
N.C.,Raleigh or USA,N.J.,Brunswick |
| [Ref.: #44974] |
Country |
USA |
| [Ref.: #44974] |
Country ISO 3 Code |
USA |
| [Ref.: #44974] |
Continent |
North America |
| |
| [Ref.: #67770] |
Sample type/isolated from |
Human skin |
| |
| [Ref.: #119771] |
Sample type/isolated from |
Human, Skin |
|
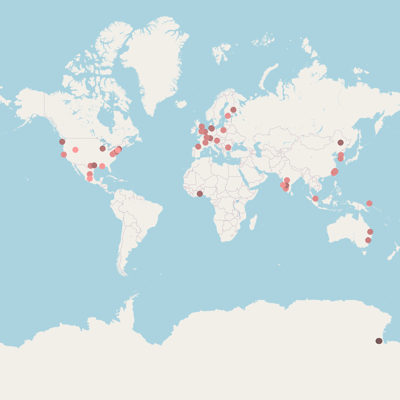
* marker position based on {}
|
|
Isolation sources categories |
| #Host |
#Human |
- |
| #Host Body-Site |
#Organ |
#Skin, Nail, Hair |
|
 |
-
Availability in culture collections
 External links External links
| [Ref.: #8672] |
Culture collection no. |
DSM 20263, ATCC 29970, CCM 2737, CCUG 7323, NCTC 11042, JCM 2416, BCRC 12923, CECT 4900, CIP 81.56, KCTC 3341, LMG 13349, NBRC 109768, NCIMB 14489, NRRL B-14755, VTT E-97770, GIFU 9124 |
| [Ref.: #83675] |
 SI-ID 7697 SI-ID 7697
|
* |
 |
Literature: |
|
Topic |
Title |
Authors |
Journal |
DOI |
Year |
|
| Pathogenicity |
Application of pulse-modulated radio-frequency atmospheric pressure glow discharge for degradation of doxycycline from a flowing liquid solution. |
Dzimitrowicz A, Caban M, Terefinko D, Pohl P, Jamroz P, Babinska W, Cyganowski P, Stepnowski P, Lojkowska E, Sledz W, Motyka-Pomagruk A |
Sci Rep |
10.1038/s41598-022-11088-w |
2022 |
* |
| Pathogenicity |
Characterization of two novel variants of staphylococcal cassette chromosome mec elements in oxacillin-resistant Staphylococcus lugdunensis. |
Chang SC, Lee MH, Yeh CF, Liu TP, Lin JF, Ho CM, Lu JJ |
J Antimicrob Chemother |
10.1093/jac/dkx291 |
2017 |
* |
| Metabolism |
5alpha-Androst-16-en-3alpha-ol beta-D-glucuronide, precursor of 5alpha-androst-16-en-3alpha-ol in human sweat. |
Starkenmann C, Mayenzet F, Brauchli R, Troccaz M |
Chem Biodivers |
10.1002/cbdv.201300286 |
2013 |
* |
|
 References References-
| #8672 |
Leibniz Institut DSMZ-Deutsche Sammlung von Mikroorganismen und Zellkulturen GmbH ; Curators of the DSMZ;
DSM 20263
|
-
-
-
| #37620 |
; Curators of the CIP;
|
-
| #44974 |
Culture Collection University of Gothenburg (CCUG) ; Curators of the CCUG;
CCUG 7323
|
-
-
| #67770 |
Japan Collection of Microorganism (JCM) ; Curators of the JCM;
|
-
| #68371 |
Automatically annotated from API 50CH acid .
|
-
| #68375 |
Automatically annotated from API ID32STA .
|
-
| #68382 |
Automatically annotated from API zym .
|
-
| #69479 |
João F Matias Rodrigues, Janko Tackmann,Gregor Rot, Thomas SB Schmidt, Lukas Malfertheiner, Mihai Danaila,Marija Dmitrijeva, Daniela Gaio, Nicolas Näpflin and Christian von Mering. University of Zurich.:
MicrobeAtlas 1.0 beta
.
|
-
-
| #83675 |
Reimer, L.C., Lissin, A.,Schober, I., Witte,J.F., Podstawka, A., Lüken, H., Bunk, B.,Overmann, J.:
StrainInfo: A central database for resolving microbial strain identifiers
.
(
DOI 10.60712/SI-ID7697.1 )
|
-
| #119771 |
Collection of Institut Pasteur ; Curators of the CIP;
CIP 81.56
|
- * These data were automatically processed and therefore are not curated
Change proposal
Successfully sent
|
|
Name and taxonomic classification

Morphology

Culture and growth conditions

Physiology and metabolism

Isolation, sampling and environmental information

Safety information

Sequence information

Genome-based predictions

External links

References