| [Ref.: #548] |
Sample type/isolated from |
hospital respirator |
| [Ref.: #548] |
Geographic location (country and/or sea, region) |
London |
| [Ref.: #548] |
Country |
United Kingdom |
| [Ref.: #548] |
Country ISO 3 Code |
GBR |
| [Ref.: #548] |
Continent |
Europe |
| |
| [Ref.: #67770] |
Sample type/isolated from |
Hospital respirator |
| |
| [Ref.: #121633] |
Sample type/isolated from |
Human, Hospital respirator |
| [Ref.: #121633] |
Geographic location (country and/or sea, region) |
London |
| [Ref.: #121633] |
Country |
United Kingdom |
| [Ref.: #121633] |
Country ISO 3 Code |
GBR |
| [Ref.: #121633] |
Continent |
Europe |
| [Ref.: #121633] |
Isolation date |
1969 |
|
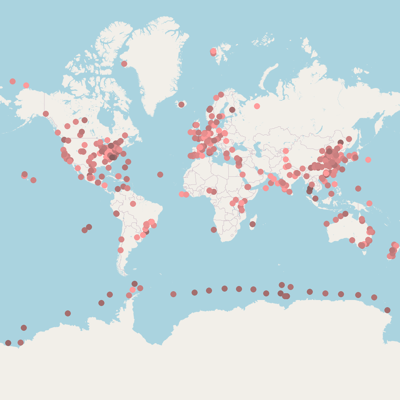
* marker position based on {}
|
|
Isolation sources categories |
| #Infection |
#Medical device |
- |
| #Infection |
#Medical environment |
#Clinic |
|
 |
-
Information on genomic background e.g. entries in nucleic sequence databass
 Sequence information Sequence information
|
16S Sequence information: |
Only first 5 entries are displayed. Click here to see all.Click here to see only first 5 entries. |
|
|
Sequence accession description |
Seq. accession number |
Sequence length (bp) |
Sequence database |
Associated NCBI tax ID |
|
| [Ref.: #548] |
Sphingomonas paucimobilis partial 16S rRNA gene, type strain DSM 1098T |
LN681566 |
1441 |
|
13689 tax ID tax ID |
| [Ref.: #67770] |
Sphingomonas paucimobilis JCM 7516 gene for 16S ribosomal RNA, partial sequence |
LC504030 |
1411 |
|
13689 tax ID tax ID |
| [Ref.: #67770] |
Sphingomonas paucimobilis gene for 16S rRNA, partial sequence |
D16144 |
1446 |
|
13689 tax ID tax ID |
| [Ref.: #67770] |
Sphingomonas paucimobilis gene for 16S ribosomal RNA, partial sequence, strain: JCM 7516 |
LC069038 |
1411 |
|
13689 tax ID tax ID |
| [Ref.: #67770] |
Sphingomonas paucimobilis gene for 16S rRNA, partial sequence, strain: NBRC 13935 |
AB680526 |
1414 |
|
13689 tax ID tax ID |
| [Ref.: #20218] |
Sphingomonas paucimobilis 16S small subunit ribosomal RNA gene, partial sequence |
U20776 |
1231 |
|
13689 tax ID tax ID |
* |
| [Ref.: #20218] |
Sphingomonas paucimobilis strain ATCC 29837 16S ribosomal RNA (rrn) gene, partial sequence |
U37337 |
1414 |
|
13689 tax ID tax ID |
* |
| [Ref.: #20218] |
Sphingomonas paucimobilis partial 16S rRNA gene, isolate OS-64.a |
AM237364 |
1431 |
|
13689 tax ID tax ID |
* |
|
| [Ref.: #548] |
GC-content |
65.0 mol% |
| [Ref.: #548] |
GC-content |
65.5 mol% |
thermal denaturation, midpoint method (Tm) |
| [Ref.: #67770] |
GC-content |
63.7 mol% |
high performance liquid chromatography (HPLC) |
|
-
Availability in culture collections
 External links External links
| [Ref.: #548] |
Culture collection no. |
DSM 1098, ATCC 29837, NCPPB 3838, NCTC 11030, JCM 7516, CCM 3293, CCUG 31192, CCUG 6518, CDC CL 1/70, CECT 599, CIP 100752, GIFU 2395, IAM 12576, ICPB 4235, IFO 13935, LMG 1227, NBRC 13935, PCM 2585 |
| [Ref.: #83360] |
 SI-ID 72295 SI-ID 72295
|
* |
 |
Literature: |
Only first 10 entries are displayed. Click here to see all.Click here to see only first 10 entries. |
|
 |
Title |
Authors |
Journal |
DOI |
Year |
|
| Phylogeny |
Sphingomonas aeria sp. nov. from indoor air of a pharmaceutical environment. |
Park HK, Han JH, Kim TS, Joung Y, Cho SH, Kwon SW, Kim SB |
Antonie Van Leeuwenhoek |
10.1007/s10482-014-0302-5 |
2014 |
* |
| Phylogeny |
Sphingomonas starnbergensis sp. nov., isolated from a prealpine freshwater lake. |
Chen H, Jogler M, Tindall BJ, Klenk HP, Rohde M, Busse HJ, Overmann J |
Int J Syst Evol Microbiol |
10.1099/ijs.0.042887-0 |
2012 |
* |
| Phylogeny |
Sphingomonas alpina sp. nov., a psychrophilic bacterium isolated from alpine soil. |
Margesin R, Zhang DC, Busse HJ |
Int J Syst Evol Microbiol |
10.1099/ijs.0.035964-0 |
2011 |
* |
| Phylogeny |
Hexachlorocyclohexane-degrading bacterial strains Sphingomonas paucimobilis B90A, UT26 and Sp+, having similar lin genes, represent three distinct species, Sphingobium indicum sp. nov., Sphingobium japonicum sp. nov. and Sphingobium francense sp. nov., and reclassification of [Sphingomonas] chungbukensis as Sphingobium chungbukense comb. nov. |
Pal R, Bala S, Dadhwal M, Kumar M, Dhingra G, Prakash O, Prabagaran SR, Shivaji S, Cullum J, Holliger C, Lal R |
Int J Syst Evol Microbiol |
10.1099/ijs.0.63201-0 |
2005 |
* |
| Enzymology |
Complement activation by bacterial surface glycolipids: a study with planar bilayer membranes. |
Munstermann M, Wiese A, Brandenburg K, Zahringer U, Brade L, Kawahara K, Seydel U |
J Membr Biol |
10.1007/s002329900486 |
1999 |
* |
| Metabolism |
Molecular mechanisms of polymyxin B-membrane interactions: direct correlation between surface charge density and self-promoted transport. |
Wiese A, Munstermann M, Gutsmann T, Lindner B, Kawahara K, Zahringer U, Seydel U |
J Membr Biol |
10.1007/s002329900350 |
1998 |
* |
| Phylogeny |
Aromatic-degrading Sphingomonas isolates from the deep subsurface. |
Fredrickson JK, Balkwill DL, Drake GR, Romine MF, Ringelberg DB, White DC |
Appl Environ Microbiol |
10.1128/aem.61.5.1917-1922.1995 |
1995 |
* |
| Phenotype |
[Phenotype characteristics of 63 strains of Pseudomonas paucimobilis, mostly of hospital origin]. |
Richard C |
Ann Biol Clin (Paris) |
|
1986 |
* |
| Phylogeny |
The similarities between Pseudomonas paucimobilis and allied bacteria derived from analysis of deoxyribonucleic acids and electrophoretic protein patterns. |
Owen RJ, Jackman PJ |
J Gen Microbiol |
10.1099/00221287-128-12-2945 |
1982 |
* |
| Phylogeny |
Isolation of an unusual 'lipid A' type glycolipid from Pseudomonas paucimobilis. |
Kawahara K, Uchida K, Aida K |
Biochim Biophys Acta |
10.1016/0005-2760(82)90285-5 |
1982 |
* |
|
Characterization of the Aerobic Anoxygenic Phototrophic Bacterium Sphingomonas sp. AAP5. |
Kopejtka K, Zeng Y, Kaftan D, Selyanin V, Gardian Z, Tomasch J, Sommaruga R, Koblizek M |
Microorganisms |
10.3390/microorganisms9040768 |
2021 |
* |
|
Enantioselective Resolution of (R,S)-Carvedilol to (S)-(-)-Carvedilol by Biocatalysts. |
Ettireddy S, Chandupatla V, Veeresham C |
Nat Prod Bioprospect |
10.1007/s13659-016-0118-2 |
2017 |
* |
|
 References References-
| #548 |
Leibniz Institut DSMZ-Deutsche Sammlung von Mikroorganismen und Zellkulturen GmbH ; Curators of the DSMZ;
DSM 1098
|
-
-
-
| #36454 |
; Curators of the CIP;
|
-
-
| #67770 |
Japan Collection of Microorganism (JCM) ; Curators of the JCM;
|
-
| #68369 |
Automatically annotated from API 20NE .
|
-
| #68382 |
Automatically annotated from API zym .
|
-
| #69479 |
João F Matias Rodrigues, Janko Tackmann,Gregor Rot, Thomas SB Schmidt, Lukas Malfertheiner, Mihai Danaila,Marija Dmitrijeva, Daniela Gaio, Nicolas Näpflin and Christian von Mering. University of Zurich.:
MicrobeAtlas 1.0 beta
.
|
-
-
-
| #83360 |
Reimer, L.C., Lissin, A.,Schober, I., Witte,J.F., Podstawka, A., Lüken, H., Bunk, B.,Overmann, J.:
StrainInfo: A central database for resolving microbial strain identifiers
.
(
DOI 10.60712/SI-ID72295.1 )
|
-
| #121633 |
Collection of Institut Pasteur ; Curators of the CIP;
CIP 100752
|
- * These data were automatically processed and therefore are not curated
Change proposal
Successfully sent
|
|
Name and taxonomic classification

Morphology

Culture and growth conditions

Physiology and metabolism

Isolation, sampling and environmental information

Safety information

Sequence information

Genome-based predictions

External links

References