-
Availability in culture collections
 External links External links
| [Ref.: #9158] |
Culture collection no. |
DSM 30137, ATCC 14482, JCM 20683, NBRC 14785, NRRL L-321, VKM B-1966, IAM 12612, CIP 107329, LMG 8819, ORS 662 |
| [Ref.: #82796] |
 SI-ID 265209 SI-ID 265209
|
* |
 |
Literature: |
|
Topic |
Title |
Authors |
Journal |
DOI |
Year |
|
| Phylogeny |
Rhizobium sophorae sp. nov. and Rhizobium sophoriradicis sp. nov., nitrogen-fixing rhizobial symbionts of the medicinal legume Sophora flavescens. |
Jiao YS, Yan H, Ji ZJ, Liu YH, Sui XH, Wang ET, Guo BL, Chen WX, Chen WF |
Int J Syst Evol Microbiol |
10.1099/ijs.0.068916-0 |
2014 |
* |
| Phylogeny |
Rhizobium etli taxonomy revised with novel genomic data and analyses. |
Lopez-Guerrero MG, Ormeno-Orrillo E, Velazquez E, Rogel MA, Acosta JL, Gonzalez V, Martinez J, Martinez-Romero E |
Syst Appl Microbiol |
10.1016/j.syapm.2012.06.009 |
2012 |
* |
| Phylogeny |
Revision of the taxonomic status of the species Rhizobium leguminosarum (Frank 1879) Frank 1889AL, Rhizobium phaseoli Dangeard 1926AL and Rhizobium trifolii Dangeard 1926AL. R. trifolii is a later synonym of R. leguminosarum. Reclassification of the strain R. leguminosarum DSM 30132 (=NCIMB 11478) as Rhizobium pisi sp. nov. |
Ramirez-Bahena MH, Garcia-Fraile P, Peix A, Valverde A, Rivas R, Igual JM, Mateos PF, Martinez-Molina E, Velazquez E |
Int J Syst Evol Microbiol |
10.1099/ijs.0.65621-0 |
2008 |
* |
|
 References References-
| #9158 |
Leibniz Institut DSMZ-Deutsche Sammlung von Mikroorganismen und Zellkulturen GmbH ; Curators of the DSMZ;
DSM 30137
|
-
-
-
| #38201 |
; Curators of the CIP;
|
-
-
| #67770 |
Japan Collection of Microorganism (JCM) ; Curators of the JCM;
|
-
| #68369 |
Automatically annotated from API 20NE .
|
-
| #68371 |
Automatically annotated from API 50CH acid .
|
-
| #68382 |
Automatically annotated from API zym .
|
-
| #69479 |
João F Matias Rodrigues, Janko Tackmann,Gregor Rot, Thomas SB Schmidt, Lukas Malfertheiner, Mihai Danaila,Marija Dmitrijeva, Daniela Gaio, Nicolas Näpflin and Christian von Mering. University of Zurich.:
MicrobeAtlas 1.0 beta
.
|
-
-
| #82796 |
Reimer, L.C., Lissin, A.,Schober, I., Witte,J.F., Podstawka, A., Lüken, H., Bunk, B.,Overmann, J.:
StrainInfo: A central database for resolving microbial strain identifiers
.
(
DOI 10.60712/SI-ID265209.1 )
|
-
| #121793 |
Collection of Institut Pasteur ; Curators of the CIP;
CIP 107329
|
- * These data were automatically processed and therefore are not curated
Change proposal
Successfully sent
|
|
Name and taxonomic classification

Morphology

Culture and growth conditions

Physiology and metabolism

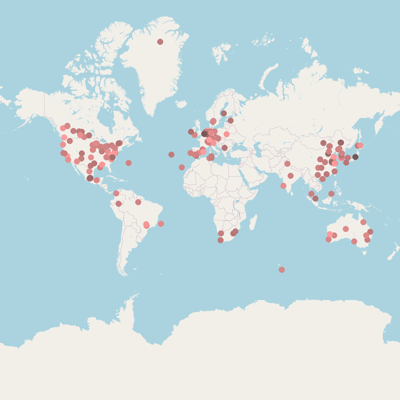
Isolation, sampling and environmental information

Safety information

Sequence information

Genome-based predictions

External links

References