| [Ref.: #34391] |
Culture collection no. |
CIP 103598, ATCC 314, JCM 2121 |
| [Ref.: #92681] |
 SI-ID 3196 SI-ID 3196
|
* |
 |
Literature: |
Only first 10 entries are displayed. Click here to see all.Click here to see only first 10 entries. |
|
Topic |
Title |
Authors |
Journal |
DOI |
Year |
|
| Enzymology |
Inhibition of Insulin Degrading Enzyme and Insulin Degradation by UV-Killed Lactobacillus acidophilus. |
Neyazi N, Motevaseli E, Khorramizadeh MR, Mohammadi Farsani T, Nouri Z, Nasli Esfahani E, Ghahremani MH |
Med Sci (Basel) |
10.3390/medsci6020036 |
2018 |
* |
| Pathogenicity |
Design and validation of an orally administrated active L. fermentum-L. acidophilus probiotic formulation using colorectal cancer Apc (Min/+) mouse model. |
Kahouli I, Malhotra M, Westfall S, Alaoui-Jamali MA, Prakash S |
Appl Microbiol Biotechnol |
10.1007/s00253-016-7885-x |
2016 |
* |
| Metabolism |
Cholesterol assimilation by Lactobacillus probiotic bacteria: an in vitro investigation. |
Tomaro-Duchesneau C, Jones ML, Shah D, Jain P, Saha S, Prakash S |
Biomed Res Int |
10.1155/2014/380316 |
2014 |
* |
| Metabolism |
Use of Lactobacillus acidophilus and Lactobacillus casei for a potential probiotic legume-based fermented product using pigeon pea (Cajanus cajan). |
Parra K, Ferrer M, Pinero M, Barboza Y, Medina LM |
J Food Prot |
10.4315/0362-028X.JFP-12-138 |
2013 |
* |
| Metabolism |
Removal of cholesterol by lactobacilli via incorporation and conversion to coprostanol. |
Lye HS, Rusul G, Liong MT |
J Dairy Sci |
10.3168/jds.2009-2574 |
2010 |
* |
| Metabolism |
Viability and growth characteristics of Lactobacillus in soymilk supplemented with B-vitamins. |
Ewe JA, Wan-Abdullah WN, Liong MT |
Int J Food Sci Nutr |
10.3109/09637480903334163 |
2010 |
* |
|
 References References-
-
| #34391 |
Collection of Institut Pasteur ; Curators of the CIP;
CIP 103598
|
-
| #67770 |
Japan Collection of Microorganism (JCM) ; Curators of the JCM;
|
-
| #68371 |
Automatically annotated from API 50CH acid .
|
-
| #68382 |
Automatically annotated from API zym .
|
-
| #69479 |
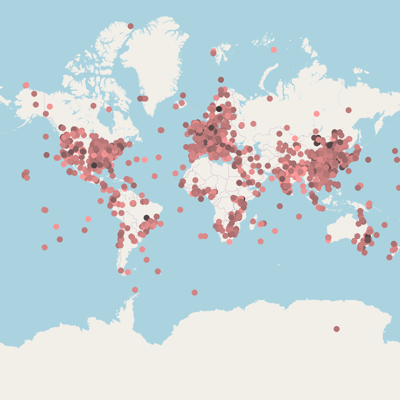
João F Matias Rodrigues, Janko Tackmann,Gregor Rot, Thomas SB Schmidt, Lukas Malfertheiner, Mihai Danaila,Marija Dmitrijeva, Daniela Gaio, Nicolas Näpflin and Christian von Mering. University of Zurich.:
MicrobeAtlas 1.0 beta
.
|
-
| #92681 |
Reimer, L.C., Lissin, A.,Schober, I., Witte,J.F., Podstawka, A., Lüken, H., Bunk, B.,Overmann, J.:
StrainInfo: A central database for resolving microbial strain identifiers
.
(
DOI 10.60712/SI-ID3196.1 )
|
- * These data were automatically processed and therefore are not curated
Change proposal
Successfully sent
|
Name and taxonomic classification

Morphology

Culture and growth conditions

Physiology and metabolism

Isolation, sampling and environmental information

Safety information

Sequence information

External links

References