-
Information on genomic background e.g. entries in nucleic sequence databass
 Sequence information Sequence information
|
16S Sequence information: |
Only first 5 entries are displayed. Click here to see all.Click here to see only first 5 entries. |
|
|
Sequence accession description |
Seq. accession number |
Sequence length (bp) |
Sequence database |
Associated NCBI tax ID |
|
| [Ref.: #20218] |
Actinosynnema mirum 16S ribosomal RNA gene, partial sequence |
AF328679 |
1509 |
|
446462 tax ID tax ID |
* |
| [Ref.: #20218] |
Actinosynnema mirum gene for 16S ribosomal RNA, partial sequence |
D85475 |
1466 |
|
40567 tax ID tax ID |
* |
| [Ref.: #20218] |
Actinosynnema mirum 16S rRNA gene, strain DSM 43827T |
X84447 |
1460 |
|
446462 tax ID tax ID |
* |
| [Ref.: #20218] |
Actinosynnema mirum clone 1 16S ribosomal RNA gene, partial sequence; 16S-23S ribosomal RNA intergenic spacer, complete sequence; and 23S ribosomal RNA gene, partial sequence |
DQ459055 |
418 |
|
446462 tax ID tax ID |
* |
| [Ref.: #20218] |
Actinosynnema mirum clone 2 16S ribosomal RNA gene, partial sequence; 16S-23S ribosomal RNA intergenic spacer, complete sequence; and 23S ribosomal RNA gene, partial sequence |
DQ459056 |
415 |
|
446462 tax ID tax ID |
* |
| [Ref.: #20218] |
Actinosynnema mirum clone 3 16S ribosomal RNA gene, partial sequence; 16S-23S ribosomal RNA intergenic spacer, complete sequence; and 23S ribosomal RNA gene, partial sequence |
DQ459057 |
413 |
|
446462 tax ID tax ID |
* |
| [Ref.: #20218] |
Actinosynnema mirum clone 6 16S ribosomal RNA gene, partial sequence; 16S-23S ribosomal RNA intergenic spacer, complete sequence; and 23S ribosomal RNA gene, partial sequence |
DQ459058 |
413 |
|
446462 tax ID tax ID |
* |
| [Ref.: #20218] |
Actinosynnema mirum clone 8 16S ribosomal RNA gene, partial sequence; 16S-23S ribosomal RNA intergenic spacer, complete sequence; and 23S ribosomal RNA gene, partial sequence |
DQ459059 |
414 |
|
446462 tax ID tax ID |
* |
| [Ref.: #20218] |
Actinosynnema mirum clone 9 16S ribosomal RNA gene, partial sequence; 16S-23S ribosomal RNA intergenic spacer, complete sequence; and 23S ribosomal RNA gene, partial sequence |
DQ459060 |
413 |
|
446462 tax ID tax ID |
* |
| [Ref.: #20218] |
Actinosynnema mirum clone 13 16S ribosomal RNA gene, partial sequence; 16S-23S ribosomal RNA intergenic spacer, complete sequence; and 23S ribosomal RNA gene, partial sequence |
DQ459065 |
418 |
|
446462 tax ID tax ID |
* |
|
|
Genome sequence information: |
|
|
Sequence accession description |
Seq. accession number |
Assembly level |
Sequence database |
Associated NCBI tax ID |
|
| [Ref.: #66792] |
Actinosynnema mirum DSM 43827 |
GCA_000023245 |
complete |
|
446462 tax ID tax ID |
* |
| [Ref.: #66792] |
Actinosynnema mirum DSM 43827 |
446462.6 |
complete |
|
446462 tax ID tax ID |
* |
| [Ref.: #66792] |
Actinosynnema mirum DSM 43827 |
644736323 |
draft |
|
446462 tax ID tax ID |
* |
|
|
-
Availability in culture collections
 External links External links
| [Ref.: #11298] |
Culture collection no. |
DSM 43827, ATCC 29888, IAM 14280, IFO 14064, IMET 9740, IMRU 3971, JCM 3225, KCC A-0225, NBRC 14064, BCRC 13449, IMSNU 20048, KCTC 9125, KCTC 9184, NCIMB 13271, NRRL B-12336, VKM Ac-843 |
| [Ref.: #82419] |
 SI-ID 39788 SI-ID 39788
|
* |
 |
Literature: |
Only first 10 entries are displayed. Click here to see all.Click here to see only first 10 entries. |
|
Topic |
Title |
Authors |
Journal |
DOI |
Year |
|
| Phylogeny |
Mirilactams C-E, Novel Polycyclic Macrolactams Isolated from Combined-Culture of Actinosynnema mirum NBRC 14064 and Mycolic Acid-Containing Bacterium. |
Hoshino S, Ozeki M, Wong CP, Zhang H, Hayashi F, Awakawa T, Morita H, Onaka H, Abe I |
Chem Pharm Bull (Tokyo) |
10.1248/cpb.c18-00143 |
2018 |
* |
| Biotechnology |
Metabolic engineering of Corynebacterium glutamicum for production of sunscreen shinorine. |
Tsuge Y, Kawaguchi H, Yamamoto S, Nishigami Y, Sota M, Ogino C, Kondo A |
Biosci Biotechnol Biochem |
10.1080/09168451.2018.1452602 |
2018 |
* |
| Metabolism |
Identification and characterization of two types of amino acid-regulated acetyltransferases in actinobacteria. |
Lu YX, Liu XX, Liu WB, Ye BC |
Biosci Rep |
10.1042/BSR20170157 |
2017 |
* |
| Genetics |
Genome sequencing and annotation of Amycolatopsis vancoresmycina strain DSM 44592(T). |
Kaur N, Kumar S, Mayilraj S |
Genom Data |
10.1016/j.gdata.2013.10.006 |
2013 |
* |
| Metabolism |
Discovery of gene cluster for mycosporine-like amino acid biosynthesis from Actinomycetales microorganisms and production of a novel mycosporine-like amino acid by heterologous expression. |
Miyamoto KT, Komatsu M, Ikeda H |
Appl Environ Microbiol |
10.1128/AEM.00727-14 |
2014 |
* |
| Metabolism |
Comparative metabolomic analysis of an alternative biosynthetic pathway to pseudosugars in Actinosynnema mirum DSM 43827. |
Asamizu S, Abugreen M, Mahmud T |
Chembiochem |
10.1002/cbic.201300384 |
2013 |
* |
|
Protein acetylation: an important mechanism in actinobacteria. |
Zhang H, Xu X |
Biosci Rep |
10.1042/BSR20170851 |
2018 |
* |
|
 References References-
| #11298 |
Leibniz Institut DSMZ-Deutsche Sammlung von Mikroorganismen und Zellkulturen GmbH ; Curators of the DSMZ;
DSM 43827
|
-
-
-
-
-
| #43417 |
Miriam Land, Alla Lapidus, Shanmugam Mayilraj, Feng Chen, Alex Copeland, Tijana Glavina Del Rio, Matt Nolan, Susan Lucas, Hope Tice, Jan-Fang Cheng, Olga Chertkov, David Bruce, Lynne Goodwin, Sam Pitluck, Manfred Rohde, Markus Göker, Amrita Pati, Natalia Ivanova, Konstantinos Mavromatis, Amy Chen, Krishna Palaniappan, Loren Hauser, Yun-Juan Chang, Cynthia C. Jeffries, Thomas Brettin, John C. Detter, Cliff Han, Patrick Chain, Brian J. Tindall, Jim Bristow, Jonathan A. Eisen, Victor Markowitz, Philip Hugenholtz, Nikos C. Kyrpides and Hans-Peter Klenk:
Complete genome sequence of Actinosynnema mirum type strain (101).
Stand Genomic Sci 1:
46 - 53
2009 (
DOI 10.4056/sigs.21137 , PubMed 21304636 )
|
-
-
-
| #66794 |
Antje Chang, Lisa Jeske, Sandra Ulbrich, Julia Hofmann, Julia Koblitz, Ida Schomburg, Meina Neumann-Schaal, Dieter Jahn, Dietmar Schomburg:
BRENDA, the ELIXIR core data resource in 2021: new developments and updates.
Nucleic Acids Res. 49:
D498 - D508
2020 (
DOI 10.1093/nar/gkaa1025 , PubMed 33211880 )
|
-
| #67770 |
Japan Collection of Microorganism (JCM) ; Curators of the JCM;
|
-
| #68368 |
Automatically annotated from API 20E .
|
-
| #68369 |
Automatically annotated from API 20NE .
|
-
| #68382 |
Automatically annotated from API zym .
|
-
-
-
| #82419 |
Reimer, L.C., Lissin, A.,Schober, I., Witte,J.F., Podstawka, A., Lüken, H., Bunk, B.,Overmann, J.:
StrainInfo: A central database for resolving microbial strain identifiers
.
(
DOI 10.60712/SI-ID39788.1 )
|
- * These data were automatically processed and therefore are not curated
Change proposal
Successfully sent
|
|
Name and taxonomic classification

Morphology

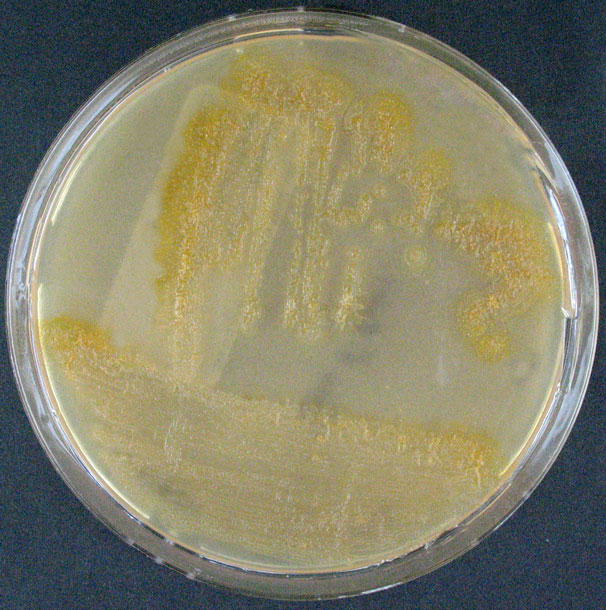
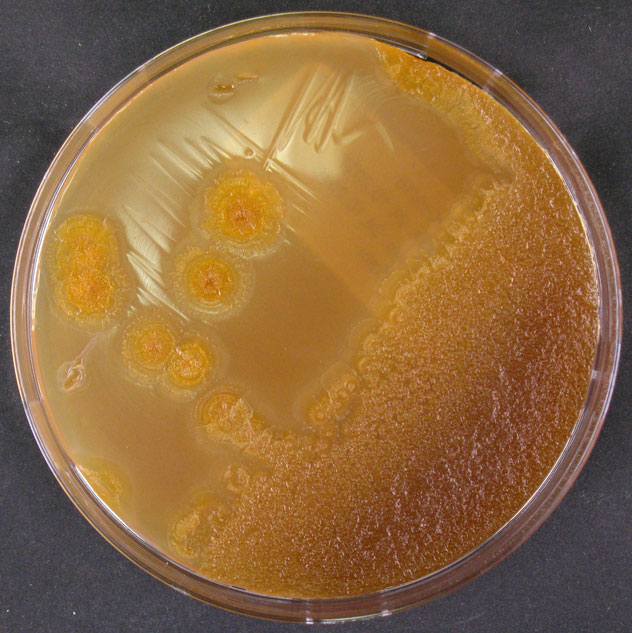
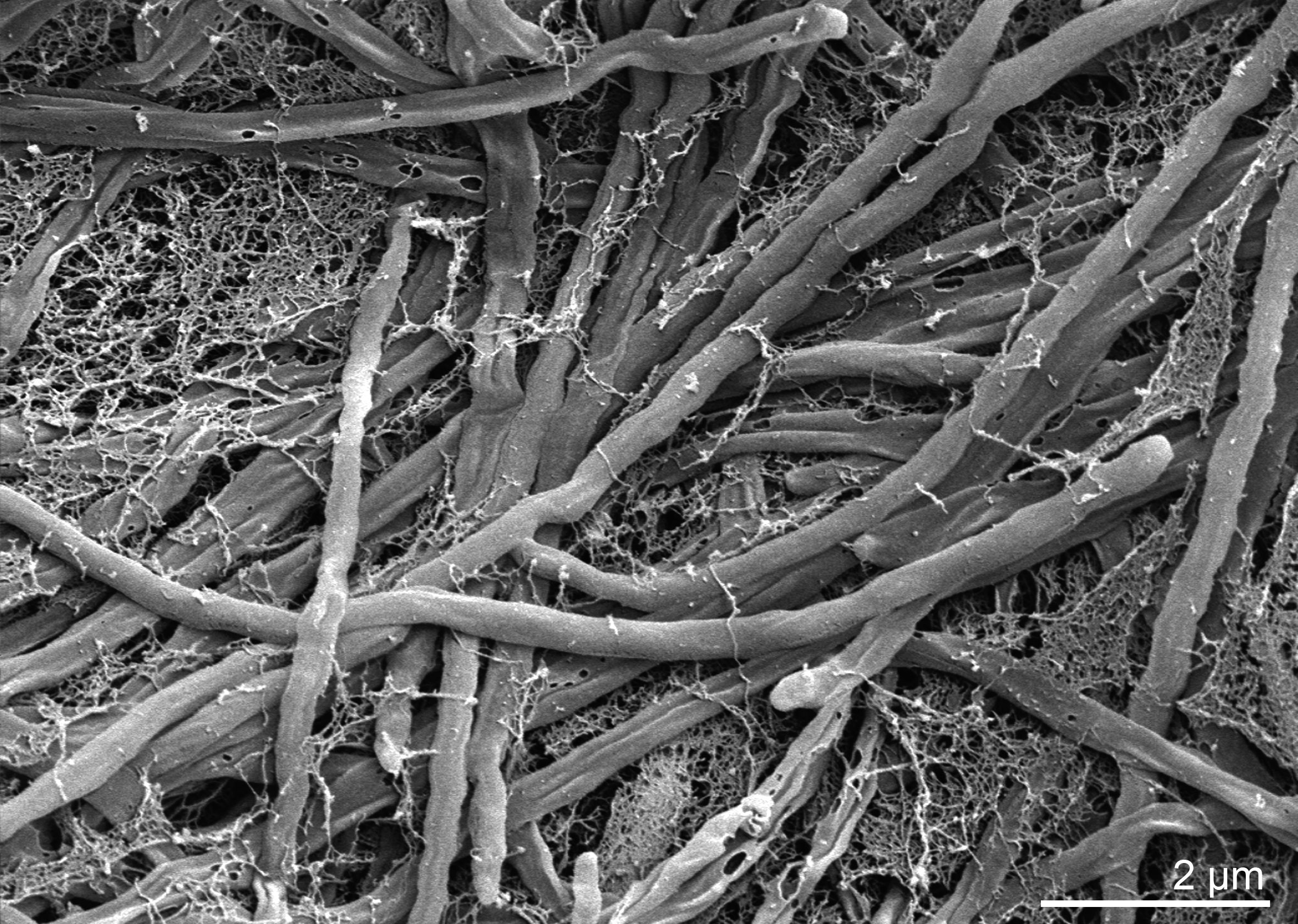
Culture and growth conditions

Physiology and metabolism

Isolation, sampling and environmental information

Safety information

Sequence information

Genome-based predictions

External links

References








