| [Ref.: #3499] |
Sample type/isolated from |
skin slime of pufferfish (Fugu poecilonotus) |
| [Ref.: #3499] |
Host species |
Fugu poecilonotus |
| |
| [Ref.: #67770] |
Sample type/isolated from |
Skin slime of pufferfish (Fugu poecilonotus) |
| [Ref.: #67770] |
Host species |
Fugu poecilonotus |
| |
| [Ref.: #123491] |
Sample type/isolated from |
Animal, Pufferfish (Fugu poecilonotus), skin slime |
| [Ref.: #123491] |
Geographic location (country and/or sea, region) |
Sendai Bay |
| [Ref.: #123491] |
Country |
Japan |
| [Ref.: #123491] |
Country ISO 3 Code |
JPN |
| [Ref.: #123491] |
Continent |
Asia |
| [Ref.: #123491] |
Isolation date |
1985-07-01 |
|
* marker position based on {}
|
|
Isolation sources categories |
| #Host |
#Fishes |
- |
| #Host Body-Site |
#Organ |
#Skin, Nail, Hair |
| #Host Body Product |
#Fluids |
- |
|
 |
-
Availability in culture collections
 External links External links
| [Ref.: #3499] |
Culture collection no. |
DSM 9166, IAM 14160, NCIMB 13177, JCM 21038, ATCC 51193, CIP 104758, KMM 458, NBRC 103034, CIP 107120 |
| [Ref.: #81898] |
 SI-ID 42852 SI-ID 42852
|
* |
 |
Literature: |
|
Topic |
Title |
Authors |
Journal |
DOI |
Year |
|
| Genetics |
Structure of a colitose-containing O-specific polysaccharide of the marine bacterium Pseudoalteromonas tetraodonis IAM 14160(T). |
Muldoon J, Perepelov AV, Shashkov AS, Gorshkova RP, Nazarenko EL, Zubkov VA, Ivanova EP, Knirel YA, Savage AV |
Carbohydr Res |
10.1016/s0008-6215(01)00121-5 |
2001 |
* |
| Phylogeny |
Retrieval of the species Alteromonas tetraodonis Simidu et al. 1990 as Pseudoalteromonas tetraodonis comb. nov. and emendation of description. |
Ivanova EP, Romanenko LA, Matte MH, Matte GR, Lysenko AM, Simidu U, Kita-Tsukamoto K, Sawabe T, Vysotskii MV, Frolova GM, Mikhailov V, Christen R, Colwell RR |
Int J Syst Evol Microbiol |
10.1099/00207713-51-3-1071 |
2001 |
* |
|
 References References-
| #3499 |
Leibniz Institut DSMZ-Deutsche Sammlung von Mikroorganismen und Zellkulturen GmbH ; Curators of the DSMZ;
DSM 9166
|
-
-
-
| #42032 |
; Curators of the CIP;
|
-
-
| #67770 |
Japan Collection of Microorganism (JCM) ; Curators of the JCM;
|
-
| #68382 |
Automatically annotated from API zym .
|
-
| #69479 |
João F Matias Rodrigues, Janko Tackmann,Gregor Rot, Thomas SB Schmidt, Lukas Malfertheiner, Mihai Danaila,Marija Dmitrijeva, Daniela Gaio, Nicolas Näpflin and Christian von Mering. University of Zurich.:
MicrobeAtlas 1.0 beta
.
|
-
-
-
| #81898 |
Reimer, L.C., Lissin, A.,Schober, I., Witte,J.F., Podstawka, A., Lüken, H., Bunk, B.,Overmann, J.:
StrainInfo: A central database for resolving microbial strain identifiers
.
(
DOI 10.60712/SI-ID42852.1 )
|
-
| #123491 |
Collection of Institut Pasteur ; Curators of the CIP;
CIP 104758
|
- * These data were automatically processed and therefore are not curated
Change proposal
Successfully sent
|
|
Name and taxonomic classification

Morphology

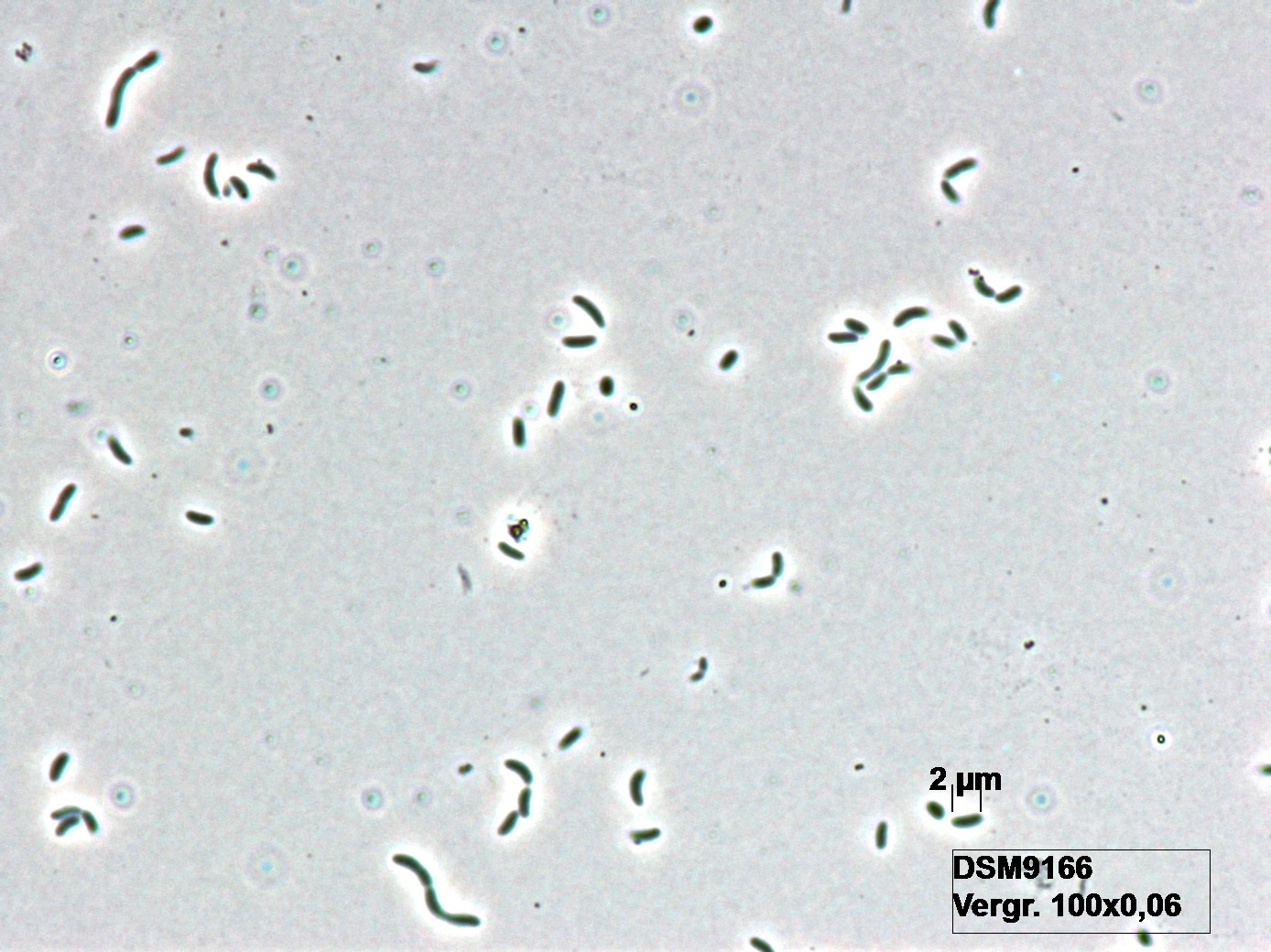
Culture and growth conditions

Physiology and metabolism

Isolation, sampling and environmental information

Safety information

Sequence information

Genome-based predictions

External links

References