| [Ref.: #3877] |
Culture collection no. |
DSM 10281, ATCC 14670, JCM 21423, CCUG 20333, LMG 2199, CIP 103296, IAM 14941, NBRC 102521, NCIMB 9462 |
| [Ref.: #80589] |
 SI-ID 4110 SI-ID 4110
|
* |
 |
Literature: |
|
Topic |
Title |
Authors |
Journal |
DOI |
Year |
|
| Phylogeny |
Reclassification of Herbaspirillum putei as a later heterotypic synonym of Herbaspirillum huttiense, with the description of H. huttiense subsp. huttiense subsp. nov. and H. huttiense subsp. putei subsp. nov., comb. nov., and description of Herbaspirillum aquaticum sp. nov. |
Dobritsa AP, Reddy MCS, Samadpour M |
Int J Syst Evol Microbiol |
10.1099/ijs.0.009381-0 |
2009 |
* |
|
 References References-
| #3877 |
Leibniz Institut DSMZ-Deutsche Sammlung von Mikroorganismen und Zellkulturen GmbH ; Curators of the DSMZ;
DSM 10281
|
-
-
| #34240 |
; Curators of the CIP;
|
-
| #47134 |
Culture Collection University of Gothenburg (CCUG) ; Curators of the CCUG;
CCUG 20333
|
-
| #67770 |
Japan Collection of Microorganism (JCM) ; Curators of the JCM;
|
-
| #69479 |
João F Matias Rodrigues, Janko Tackmann,Gregor Rot, Thomas SB Schmidt, Lukas Malfertheiner, Mihai Danaila,Marija Dmitrijeva, Daniela Gaio, Nicolas Näpflin and Christian von Mering. University of Zurich.:
MicrobeAtlas 1.0 beta
.
|
-
| #80589 |
Reimer, L.C., Lissin, A.,Schober, I., Witte,J.F., Podstawka, A., Lüken, H., Bunk, B.,Overmann, J.:
StrainInfo: A central database for resolving microbial strain identifiers
.
(
DOI 10.60712/SI-ID4110.1 )
|
-
| #119992 |
Collection of Institut Pasteur ; Curators of the CIP;
CIP 103296
|
- * These data were automatically processed and therefore are not curated
Change proposal
Successfully sent
|
Name and taxonomic classification

Morphology

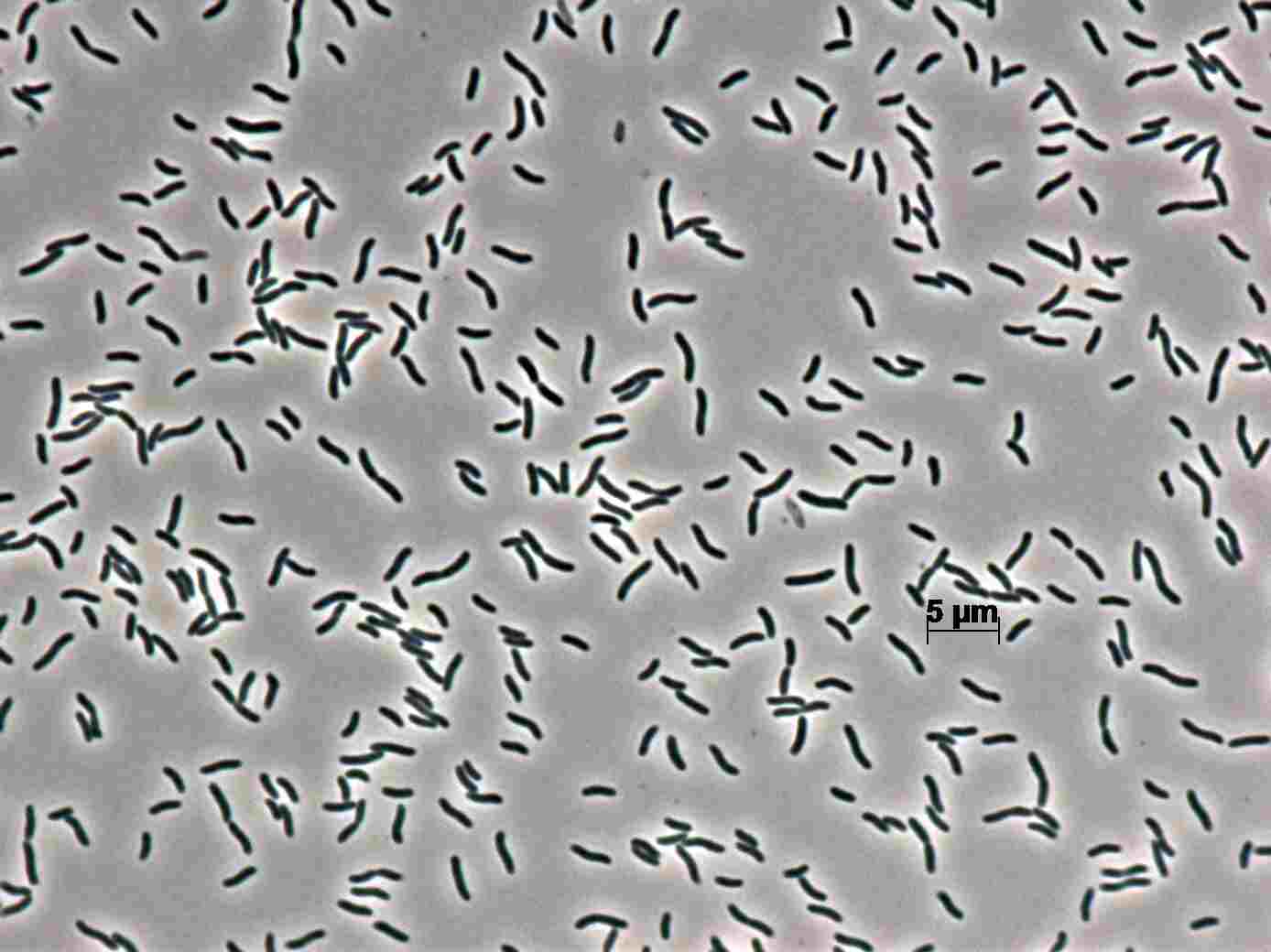
Culture and growth conditions

Physiology and metabolism

Isolation, sampling and environmental information

Safety information

Sequence information

External links

References