-
Availability in culture collections
 External links External links
| [Ref.: #9846] |
Culture collection no. |
DSM 40847, ATCC 29032, JCM 4168, KCC S-0168, BCRC 12165, CBS 199.75, CBS 207.78, CGMCC 4.1719, CGMCC 4.1851, CIP 108144, IFO 13819, NBRC 13819, NCIMB 11159, NRRL B-3729, RIA 1627, VKM Ac-928, VTT E-032076, CCRC 12165 |
| [Ref.: #85228] |
 SI-ID 361254 SI-ID 361254
|
* |
 |
Literature: |
Only first 10 entries are displayed. Click here to see all.Click here to see only first 10 entries. |
|
 |
Title |
Authors |
Journal |
DOI |
Year |
|
| Phylogeny |
Identification and Antifungal Mechanism of a Novel Actinobacterium Streptomyces huiliensis sp. nov. Against Fusarium oxysporum f. sp. cubense Tropical Race 4 of Banana. |
Qi D, Zou L, Zhou D, Zhang M, Wei Y, Zhang L, Xie J, Wang W |
Front Microbiol |
10.3389/fmicb.2021.722661 |
2021 |
* |
| Metabolism |
Features of the transglutaminase-activating metalloprotease from Streptomyces mobaraensis DSM 40847 produced in Escherichia coli. |
Juettner NE, Classen M, Colin F, Hoffmann SB, Meyners C, Pfeifer F, Fuchsbauer HL |
J Biotechnol |
10.1016/j.jbiotec.2018.07.004 |
2018 |
* |
| Genetics |
Genome Mining of Streptomyces mobaraensis DSM40847 as a Bleomycin Producer Providing a Biotechnology Platform To Engineer Designer Bleomycin Analogues. |
Hindra, Yang D, Teng Q, Dong LB, Crnovcic I, Huang T, Ge H, Shen B |
Org Lett |
10.1021/acs.orglett.7b00283 |
2017 |
* |
| Metabolism |
Involvement of a Novel Class C Beta-Lactamase in the Transglutaminase Mediated Cross-Linking Cascade of Streptomyces mobaraensis DSM 40847. |
Zindel S, Ehret V, Ehret M, Hentschel M, Witt S, Kramer A, Fiebig D, Juttner N, Frols S, Pfeifer F, Fuchsbauer HL |
PLoS One |
10.1371/journal.pone.0149145 |
2016 |
* |
| Biotechnology |
Whole-Genome Shotgun Assembly and Analysis of the Genome of Streptomyces mobaraensis DSM 40847, a Strain for Industrial Production of Microbial Transglutaminase. |
Yang H, He T, Wu W, Zhu W, Lu B, Sun W |
Genome Announc |
10.1128/genomeA.00143-13 |
2013 |
* |
| Enzymology |
Hydroxamate-based colorimetric method for direct screening of transglutaminase-producing bacteria. |
Bourneow C, Benjakul S, H-Kittikun A |
World J Microbiol Biotechnol |
10.1007/s11274-012-1017-2 |
2012 |
* |
| Metabolism |
Production of transglutaminase by Streptomyces isolates in solid-state fermentation. |
Nagy V, Szakacs G |
Lett Appl Microbiol |
10.1111/j.1472-765X.2008.02395.x |
2008 |
* |
| Enzymology |
Production of L-leucine aminopeptidase by selected Streptomyces isolates. |
Nagy V, Nampoothiri KM, Pandey A, Rahulan R, Szakacs G |
J Appl Microbiol |
10.1111/j.1365-2672.2007.03546.x |
2007 |
* |
|
Biocontrol potential and antifungal mechanism of a novel Streptomyces sichuanensis against Fusarium oxysporum f. sp. cubense tropical race 4 in vitro and in vivo. |
Qi D, Zou L, Zhou D, Zhang M, Wei Y, Li K, Zhao Y, Zhang L, Xie J |
Appl Microbiol Biotechnol |
10.1007/s00253-022-11788-3 |
2022 |
* |
|
 References References-
| #9846 |
Leibniz Institut DSMZ-Deutsche Sammlung von Mikroorganismen und Zellkulturen GmbH ; Curators of the DSMZ;
DSM 40847
|
-
-
-
-
| #36693 |
; Curators of the CIP;
|
-
-
| #66794 |
Antje Chang, Lisa Jeske, Sandra Ulbrich, Julia Hofmann, Julia Koblitz, Ida Schomburg, Meina Neumann-Schaal, Dieter Jahn, Dietmar Schomburg:
BRENDA, the ELIXIR core data resource in 2021: new developments and updates.
Nucleic Acids Res. 49:
D498 - D508
2020 (
DOI 10.1093/nar/gkaa1025 , PubMed 33211880 )
|
-
| #67770 |
Japan Collection of Microorganism (JCM) ; Curators of the JCM;
|
-
| #68368 |
Automatically annotated from API 20E .
|
-
| #68382 |
Automatically annotated from API zym .
|
-
| #69479 |
João F Matias Rodrigues, Janko Tackmann,Gregor Rot, Thomas SB Schmidt, Lukas Malfertheiner, Mihai Danaila,Marija Dmitrijeva, Daniela Gaio, Nicolas Näpflin and Christian von Mering. University of Zurich.:
MicrobeAtlas 1.0 beta
.
|
-
-
-
| #85228 |
Reimer, L.C., Lissin, A.,Schober, I., Witte,J.F., Podstawka, A., Lüken, H., Bunk, B.,Overmann, J.:
StrainInfo: A central database for resolving microbial strain identifiers
.
(
DOI 10.60712/SI-ID361254.1 )
|
-
| #120985 |
Collection of Institut Pasteur ; Curators of the CIP;
CIP 108144
|
- * These data were automatically processed and therefore are not curated
Change proposal
Successfully sent
|
|
Name and taxonomic classification

Morphology

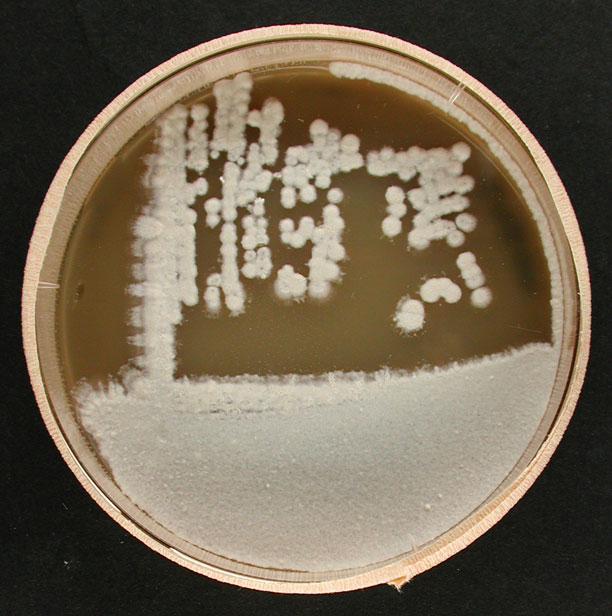
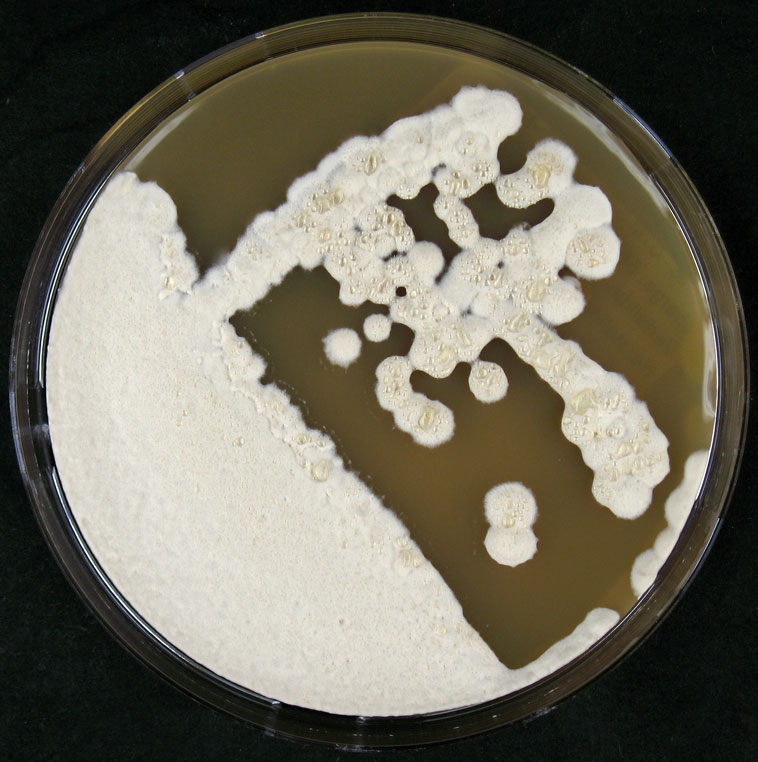
Culture and growth conditions

Physiology and metabolism

Isolation, sampling and environmental information

Safety information

Sequence information

Genome-based predictions

External links

References